The Genome
Size of a genome doesn't exessarily reflect the complexity of a species or organism as most of a genome is non-functional.
In humans only 2% codes for proteins (structural genes)
Remainder:
Promotors
Non functional = e.g. repetitive elements
Karyotype = a visual representation of an organism's chromosomes, which can be used to identify abnormalities in chromosome number or structure
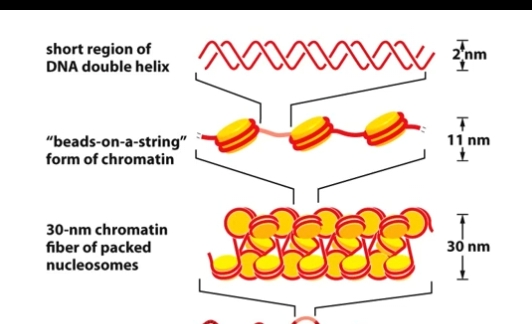
Chromatin Structure
Nucleosome = DNA wound around a histone core
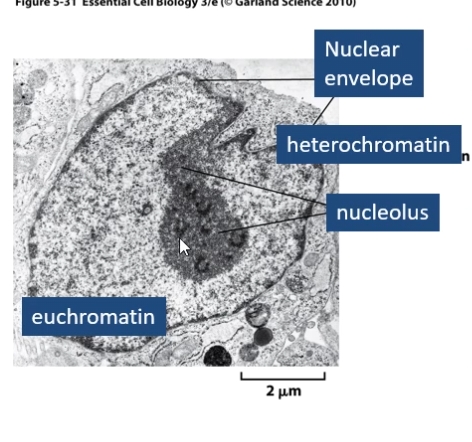
Nucleolus = structure where rRNA ia transcribed and ribosomal subunits are assembled
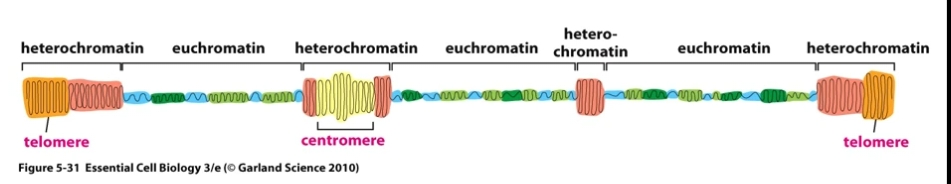
Heterochromatin = Highly condensed chromatin . Generally gene poor and transcriptionally inactive as it's inaccessible to transcription factors.
Euchromatin = Prevalent in gene-rich areas, less compact allowing access for proteins involved in transcription.
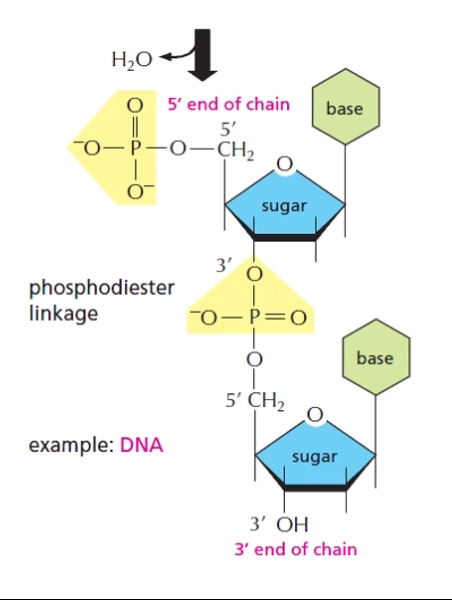
DNA structure
Remember DNA is read from 5’ end to 3’ end.
Phosphodiester linkages between 5’ phosphate group on one nucleotide and 3’ oxygen on another.
DNA Replication
Semi-conservative DNA replication = the process by which DNA is duplicated in cells to create two identical daughter strands, each containing one original strand and one new strand. Ensures that replication is accurate.
DNA polymerase synthesises strands in their 5’ to 3’ direction.
Replication bubble = an open region of DNA where the double helix unwinds and separates to allow for DNA replication. There is more than one replication bubble along a strand.
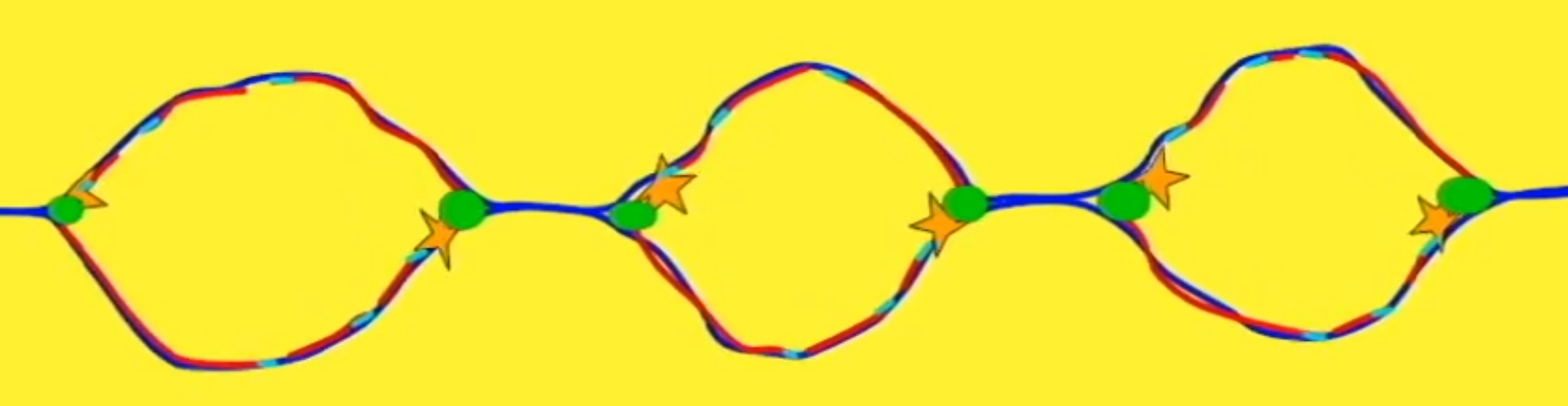
Replication origin = a specific DNA sequence where DNA replication begins on a chromosome, plasmid, or virus - TATA regions (rich in A and T bases as they have fewer H bonds between them).
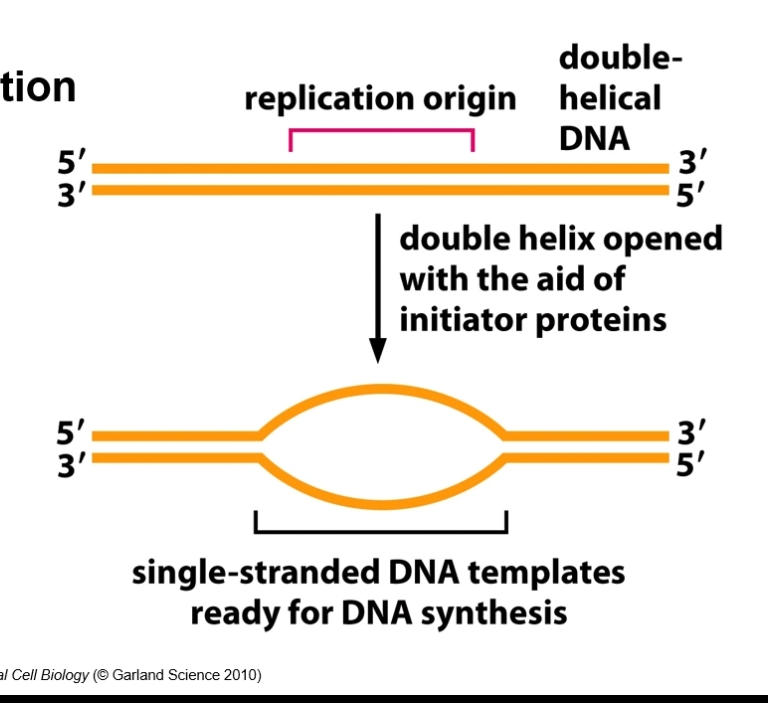
Single strand binding proteins = bind to strands in the replication bubble.
Stop strand from rejoining
Unwound ssDNA (single stranded) are susceptible to nucleases (cut DNA). Ss binding proteins protect from that enzyme
Replication fork: Y-shape structure
Replication happens in both directions in mammals = 2 DNA helicases and 2 replication forks per bubble.
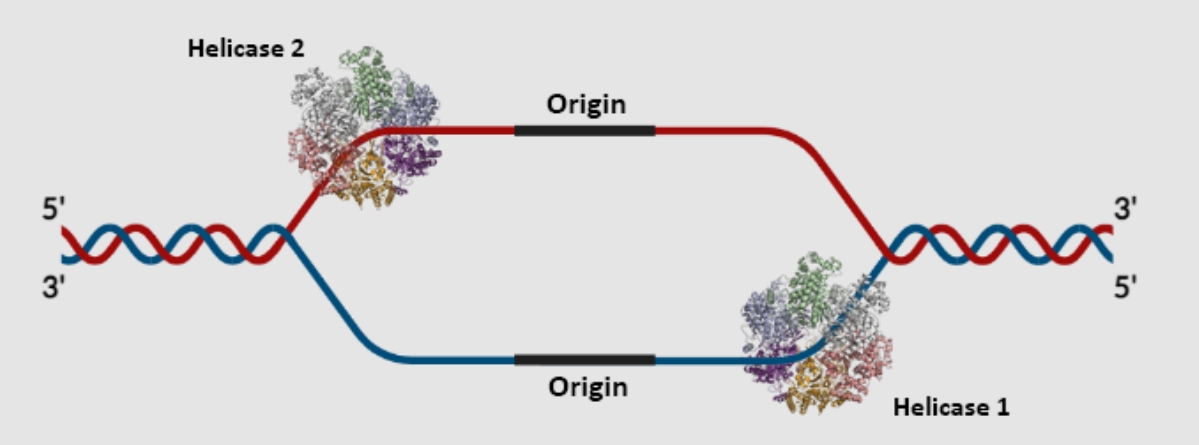
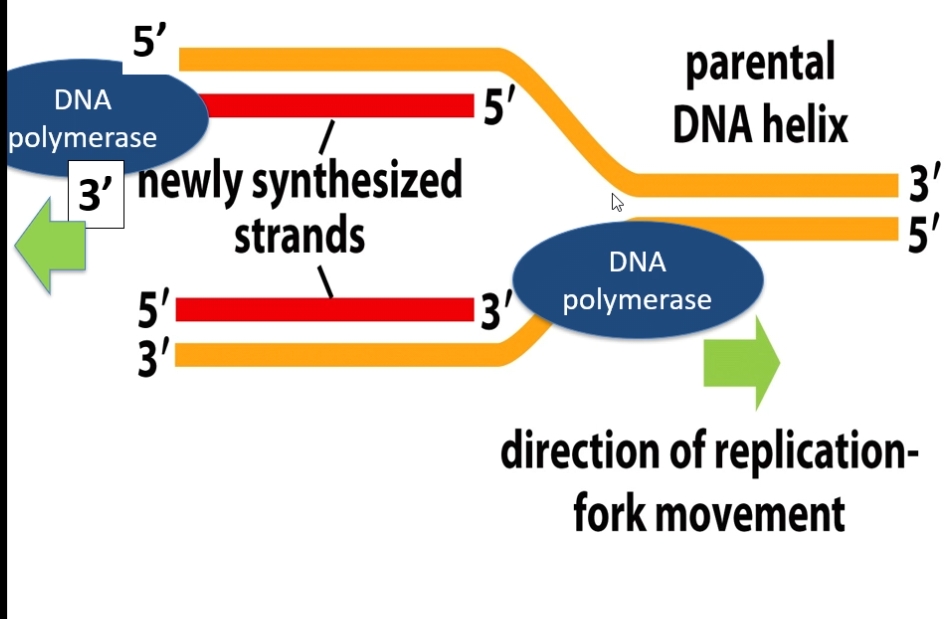
DNA polymerase on both parent strands adds nucleotides 5’ to 3’.
As DNA becomes unwound, the DNA infront of (the right side on the image above) the replication fork becomes supercoiled.
Topoisomerases (a nuclease and ligase) = introduce breaks in these regions to unwind it from being supercoiled. It also ligates the the fragmented DNA. It regulates DNA’s 3D strucure. Anti-cancer drugs could dampen the ligas activity and increase the nuclease activity of this .
Leading strand = synthesised continuously in 5’ to 3’ (same direction as replication
Lagging strand = synthesided in opposite direction, discontinuously. Made up of okazaki fragments which are later joined together by DNA ligase.
RNA primer = a short, single-stranded RNA segment complementary to a section of ss DNA template strands. These provide a starting point for DNA polymerase as polymerase can only add nucleotides to the 3’ end of a strand as it uses the OH group as a ‘hook'.
RNA primer provides the OH group for DNA polymerase. Once one nucleotide binds to primer, DNA polymerase can continue adding nucleotides as OH groups are available. On leading strand this is continuous, on lagging strand this creates many okazaki fragments.
Leading strand - only one needed on each bubble at the 5’ end (3’ end of the template strand)
Lagging strand - needed multiple on each bubbleat the ends of each okazaki fragment
Nucleases detach the RNA primer from the okazaki fragment.
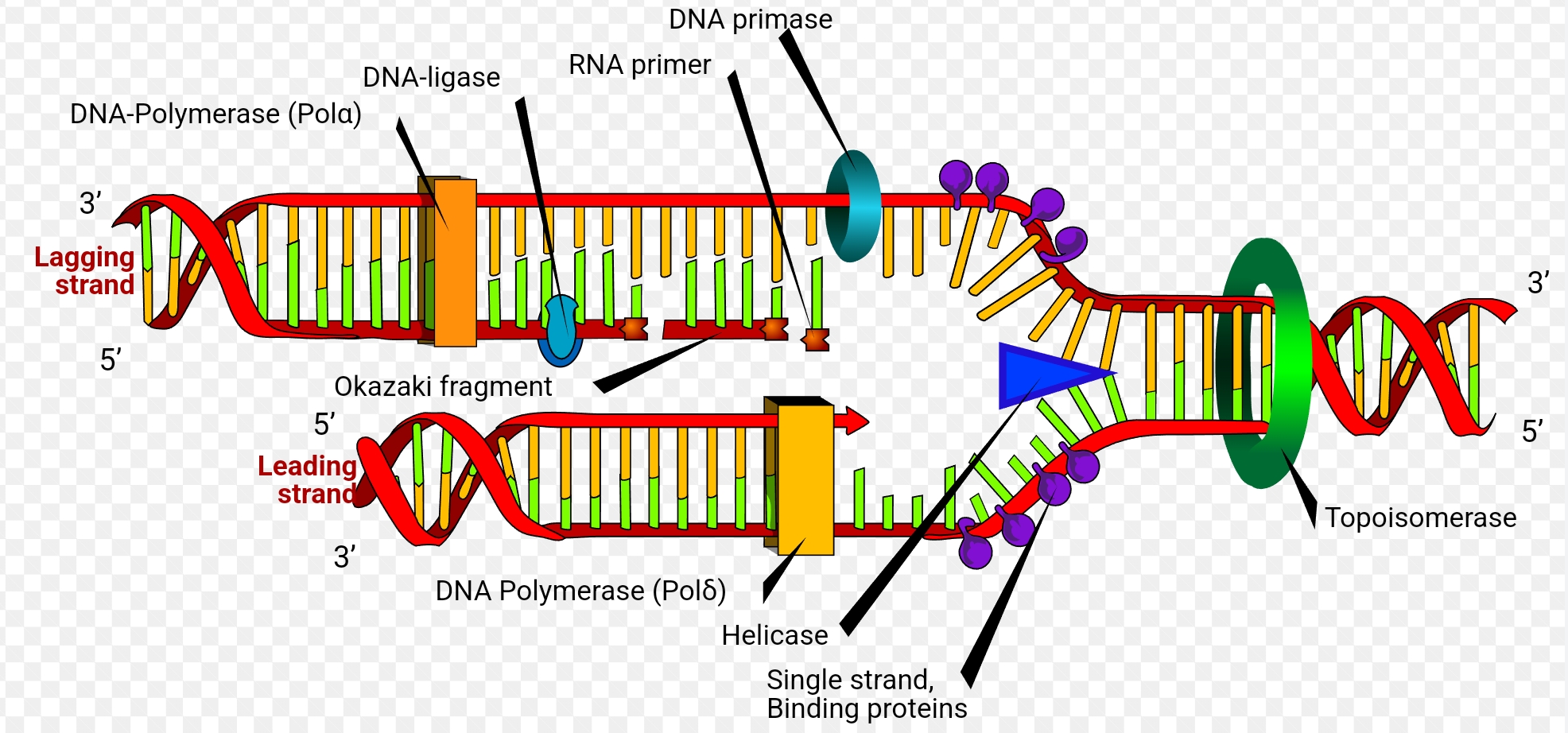
DNA ligase joins the 3 prime ends of the okazaki fragments and the 5 prime end of the partially synthesised DNA strand on lagging strand together.
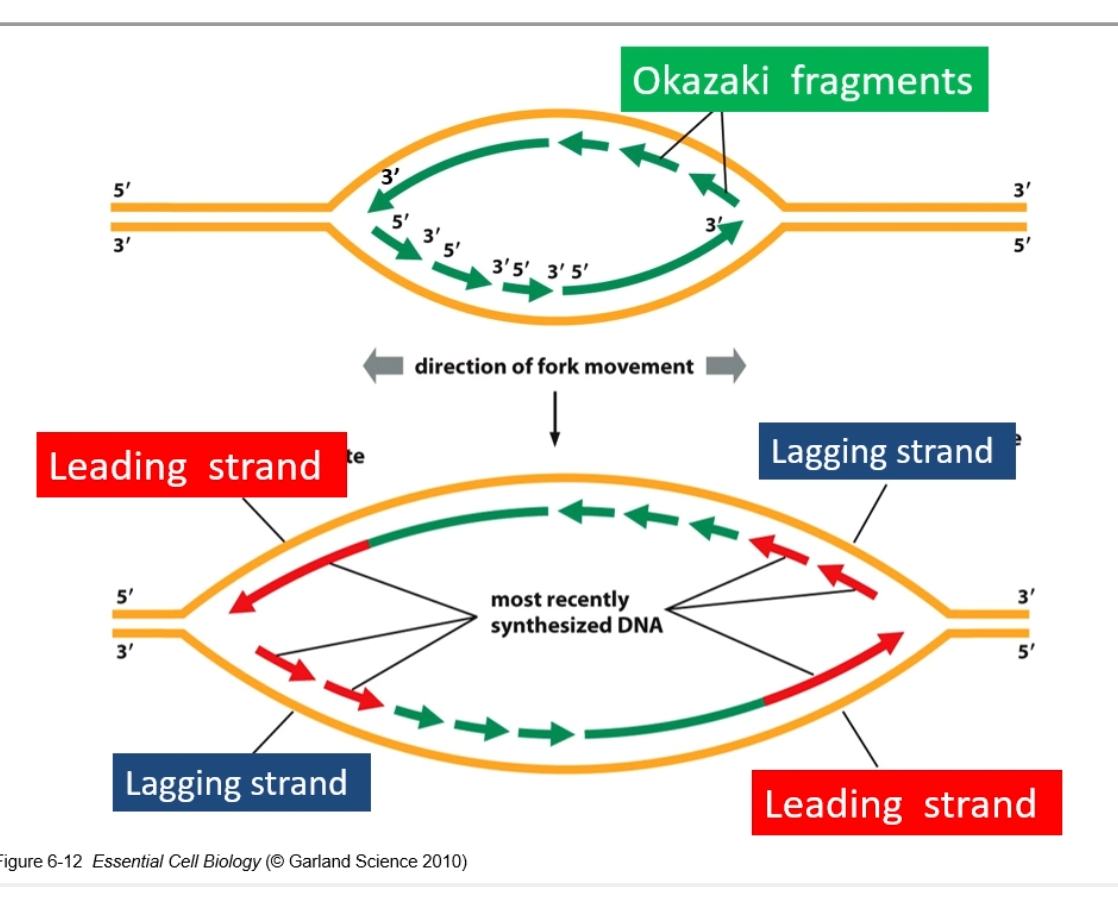
Per fork there is one lagging and leading strand. Per bubble there are 2 lagging and leading strands.
RNA primer is where there are gaps in the arrows.

DNA polymerase and proof reading
DNA polymerase makes 1 mistake per 10^7 bp
DNA P proof reads in 3’ to 5’ direction on new strand (checks the newly added base (3’ to 5’ irection) before adding the next one in the 5’ to 3’ direction)
1 mistake made in this repair system every 10^9 base pair
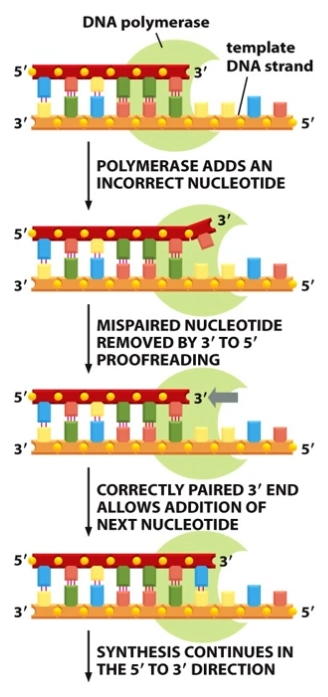
High fidelity (trustworthiness) due to:
Stability of complementary base pairing
Proof reading by DNA polymerase