Environmental Microbiology
Microbes account for around 50% of Earth’s biomass. They’re found everywhere on its surface and also deep within the Earth. We can isolate a microorganism from its environment to study its properties in lab cultures.
Isolation in pure culture is important for detailed studies in a controlled (lab) environment, as well as for applications in biotechnology eg industrial/environmental microbiology.
Microbes live in communities in the environment, making isolation in pure culture difficult.
Biofilms are assemblies of bacterial cells adhered to a surface and enclosed in an adhesive matrix excreted by the cells. The matrix is usually a mixture of polysaccharides. Biofilms trap nutrients for microbial growth and help to prevent detachment of cells in flowing systems. They also resist physical forces which sweep away unattached cells, phagocytosis by immune cells, as well as penetration of toxins eg antibiotics.
Bacteria form biofilms for several reasons including self defence, so cells can remain in a favourable niche, so cells can live in close association. Pseudomonas aeruginosa forms a biofilm in cystic fibrosis lungs. c-di-GMP is a microbial hormone which controls biofilm formation.
There are two main ways of carrying out microbial isolation from the environment: direct isolation and enrichment culture.
Direct isolation involves extracting the microbes from the sample by filtration or addition of eluent to release from solid matrix. Dilutions and plating are carried out. This is important for detailed, controlled studies and for biotech applications. Medium and incubation conditions which are selective for the desired organism are established. The conditions mimic resources and conditions in that organism’s niche. Pure cultures can be obtained in many ways eg streak plate, agar plate and liquid dilution.
For an enrichment culture, the microbes are extracted in the same way as they are for direct isolation, and enriched. They are then diluted and plated out. Enrichment culture is useful for isolation in biotech and risk assessments.
Aerobic bacteria tend to be plated for counting whereas the agar shake-tube method or an anaerobic jar tend to be used for anaerobic bacteria. Agar shake-tube method = dilution of mixed cultures in tubes of molten agar.
Direct counting may be used to count bacteria. Uses a grid. Cover slip over microscope slide means there’s a known volume under the cover slip. However live and dead cells can’t be distinguished without use of specific stains. Small cells may be overlooked and precision is difficult to achieve. If no stain is used, a phase-contrast microscope is required. Cell suspensions of low density (<10^6 cells/ml) are hard to count. Motile cells must be immobilised and debris may be mistaken. Therefore it’s likely number counted will be an overestimate.
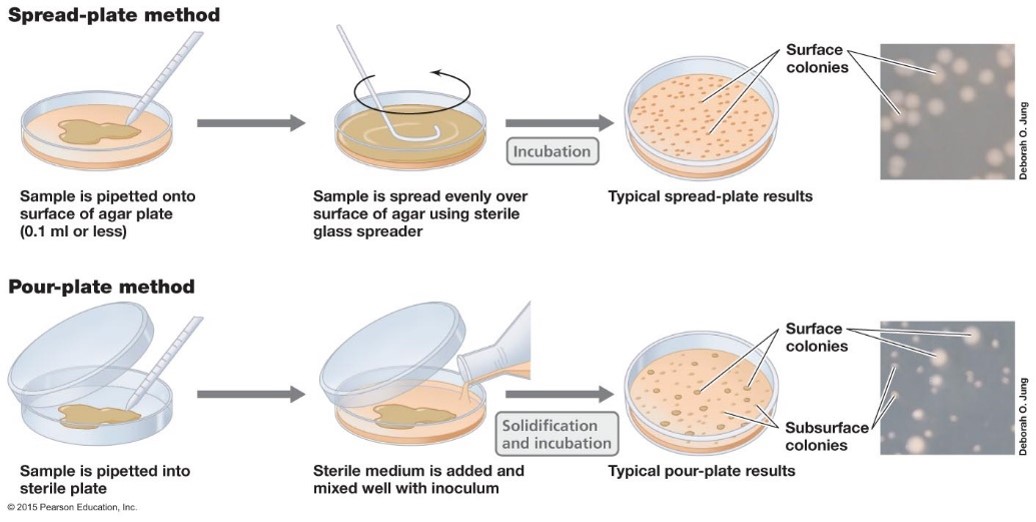
Viable cell counts = plate counts. They are a measurement of the living and reproducing population. There are two main ways to perform plate counts, the spread-plate and pour-plate method. To obtain the appropriate colony number, the sample being counted should be diluted.
Most probable numbers (MPN). This is where a serial dilution of inoculum in liquid medium is carried out until the final tube shows no growth. This is repeated several times until pure culture culture is obtained. Purity is checked by microscopy, colony characteristics on a plate, growth in other media. If bacteria grows in a media where the sample should grow poorly, this is evidence of contamination. MPNs are used for estimation of microbial numbers for routine sampling eg in food or wastewater.
Bacteria must balance their growth and death rates, either by oligotrophy or copiotrophy.
Autochthonous microbes or k strategists are oligotrophs. These are microbes which grow continuously but with low rates of activity. They have nutrient uptake enzymes with low specificity and high affinity and are adapted to low nutrient concentrations.
Zymogenous microbes or r strategists are copiotrophs. These are microbes which primarily exist in a resting phase but with brief periods of activity. Their nutrient uptake enzymes have high specificity but low affinity.
The distinction between oligotrophs and copiotrophs is often blurred eg arthrobacter are oligotrophs which undergo cell rounding and reduction in growth and Streptomyces are copiotrophs with metabolically active spores.
Viable but nonculturable bacteria (VBNC) live in a state of very low metabolic activity. They don’t divide and can’t grow on standard growth media. VBNC are morphologically smaller and have reduced nutrient transport, respiration and macromolecule synthesis. VBNC arise as a response to stress eg nutrient, temperature, osmotic, oxygen or light stress. They have the ability to become culturable once resuscitated and have significant implications for public health.
Antibiotic hunting involves streaking a suspected antibiotic producer across one side of a plate. After incubating with the hopes that antibiotic permeates the agar, the plate is cross-streaked with test organisms and incubated again. Inhibition zones are observed where antibiotic was effective against certain species. Sources of antibiotic producers may be natural (eg fungi, plants, actinobacteria) synthetic or semi-synthetic.
A lawn of indicator organism may be made and overlayed with antibiotic-soaked discs. After incubation there should be zones of inhibition of various sizes surrounding some/all of the discs. Larger radius = more effective antibiotic against indicator organism.
Testing antibiotic susceptibility by dilution methods allows the minimum inhibitory concentration (MIC) to be determined. The dilutions are inoculated with test organism. Growth (turbidity) occurs at concentrations below the MIC. Antibiotic must be effective at low concentrations to be a successful drug.
New antibiotics may be found by mining and sequencing an organism’s genome. This is because sequencing the genome allows us to convert the sequences into proteins in silico. Metabolic pathways can then be constructed and antibiotics targeting these pathways can be designed.
Microbes can be used for mining. Sulphides form insoluble minerals with metals. It’s only economically feasible to mine low-grade ores if the metal can be concentrated. Microbial activity removes copper from sulphide ores.
Microbial leaching = low grade ore dumped in a large pile (leach dump) and sulphuric acid added. Liquid which emerges from the bottom of the pile is enriched in dissolved metals and transported to a precipitation plant. Bacterial oxidation of Fe (II) ions is critical in microbial leaching as Fe (III) ions can oxidise metals in the ores.
Uranium contamination of groundwater occurs where uranium has been processed or stored. Some bacteria eg Geobacter and Shewanella are able to reduce water soluble uranium (VI) ions to insoluble uranium (IV) ions.
Microbes are able to completely degrade organic pollutants to carbon dioxide. This means prokaryotes have been used in bioremediation of several major crude oil spills eg Exxon Valdez.
Diverse bacteria, fungi and some cyanobacteria and green algae can oxidise petroleum products aerobically. Oil-oxidising activity is best with optimal temperature and inorganic nutrient concentrations. Hydrocarbon-degrading bacteria attach to oil droplets, decomposing the oil and dispersing the slick.
Xenobiotic compounds are synthetic chemicals which aren’t naturally occurring eg pesticides, polychlorinated biphenyls, munitions, dyes and chlorinated solvents. Most degrade very slowly and are common components of toxic wastes. Some can be used as carbon sources or electron donors by microbes. Burkholderia species dechlorinate the pesticide 2,4,5-T aerobically.
Plastics aren’t readily degraded by microbes. This has fuelled research into biopolymers, biodegradable alternatives to plastics. Polyhydroxyalkanoates (PHAs) are linear polyesters which bacteria eg Bacillus megatarium use as storage compounds. They are biodegradable and used in bioplastic production.
PHAs are degraded by PHA depolymerase in Alcaligenes, Rhodospirillum rubrum, B. megaterium and Pseudomonas lemoignei. PHA degradation under aerobic conditions produces carbon dioxide and water, and under anaerobic conditions produces carbon dioxide and methane.