Class 3: Protein Transport and Mitochondrial Function
What we discussed last class: transport Between Cytosol and Nucleus
Transport routes exist between the cytosol and various organelles:
Nucleus: The primary site of genetic material and transcriptional regulation.
ALS and Defective Nuclear Transport
Amyotrophic Lateral Sclerosis (ALS) is a neurodegenerative disease affecting motor neurons, leading to progressive muscle weakness and paralysis.
It is a fatal motor neuron disease, with a typical survival of 2-5 years after diagnosis.
10% of cases are familial, indicating a genetic component.
C9ORF72 expansion accounts for approximately 40% of familial cases and 5-10% of sporadic cases, making it the most common genetic cause of ALS. It’s a repeated section of the gene in the promotor, and this issue with the promotor affects the expression of the gene itself.
ALS involves defective nuclear transport, disrupting the normal flow of molecules between the nucleus and cytoplasm, impacting cellular function and survival.
Experiment shows that toxic PR peptides affect the nuclear intensity of NLS-Protein-GFP. This indicates that these peptides interfere with the import of proteins into the nucleus.
There are toxic peptide repeats that if we see in a electronic microscope, can appear as aggregates within the cytoplasm, leading to cellular dysfunction and contributing to the progression of the disease. We can see in the nucleoporins where they aggregate in the green and the peptides in the red. They stick in the center of the nucleoporins and prevent entry and exit of proteins in and out of the nucleus.
As the concentration of PR toxic peptide increases, the nuclear intensity decreases, demonstrating a dose-dependent relationship between PR peptide concentration and nuclear transport efficiency.
Nuclear Pore Complex (NPC) and Transport (Review)
The nuclear envelope facilitates extensive bidirectional traffic via NPCs, regulating the movement of molecules in and out of the nucleus to maintain cellular homeostasis and proper function.
Proteins with Nuclear Localization Signals (NLSs) are actively imported into the nucleus via importin proteins, ensuring that essential proteins reach their destination for replication, transcription, and repair.
Proteins with Nuclear Export Signals (NESs) are actively exported out of the nucleus, allowing processed mRNA, ribosomal subunits, and other molecules to reach the cytoplasm for protein synthesis and other cellular processes.
Importins and exportins shuttle between the cytosol and nucleus, acting as carrier proteins that facilitate the translocation of cargo across the nuclear envelope.
GTPase Ran provides energy and directionality for transport, regulating the binding and release of cargo from importins and exportins to ensure efficient and regulated transport.
Passive transport occurs through the center of the channel, allowing small molecules and ions to diffuse freely across the nuclear envelope, while active transport occurs at the channel periphery for larger molecules and proteins.
NLSs are not removed, and proteins are imported repeatedly after mitosis, allowing for continuous functionality and maintenance of nuclear proteins.
NUP214 Mutations and Defective Nuclear Transport
Mutations in NUP214 (a member of FG-repeat containing Nucleoporins) cause:
Neurodevelopmental regression, leading to a decline in cognitive and motor skills.
Seizures, indicating abnormal electrical activity in the brain.
Microcephaly, characterized by an abnormally small head size.
Cerebral atrophy, involving the progressive loss of brain tissue.
Fibroblasts from patients with NUP214 mutations show defects in nuclear import and export, indicating that the mutations disrupt the normal function of the NPC and impair transport processes.
NUP214 mutated cells exhibit more "central plugs" in pore channels, which may represent large cargo caught in transit, affecting NPC function. This suggests that these mutations impede the passage of molecules through the NPC, leading to transport defects.
Transport into Mitochondria
Most mitochondrial proteins are imported from the cytosol to specific sub-compartments, ensuring proper function and maintenance of the organelle.
Mitochondria Structural Features include:
Cristae: Folds of the inner membrane that increase the surface area for ATP production.
Matrix: The innermost compartment of the mitochondria, containing enzymes, DNA, and ribosomes.
Inner Membrane: Contains the electron transport chain and ATP synthase.
Outer Membrane: The outer boundary of the mitochondria.
Mitochondrial Protein Import
Translocation into the mitochondria occurs via a mitochondrial targeting sequence (MTS), which guides the protein to the appropriate location within the organelle.
Fully synthesized mitochondrial precursor proteins translocate from the cytosol post-translationally, allowing for proper folding and modification before import.
For example, the import into the matrix signal (MTS) forms an alpha helix, has a N terminal.
MTS consists of positively charged amino acids clustered on one face of an α helix and nonpolar amino acids on the opposite side, facilitating interaction with import receptors and translocation machinery.
Mitochondrial Translocators
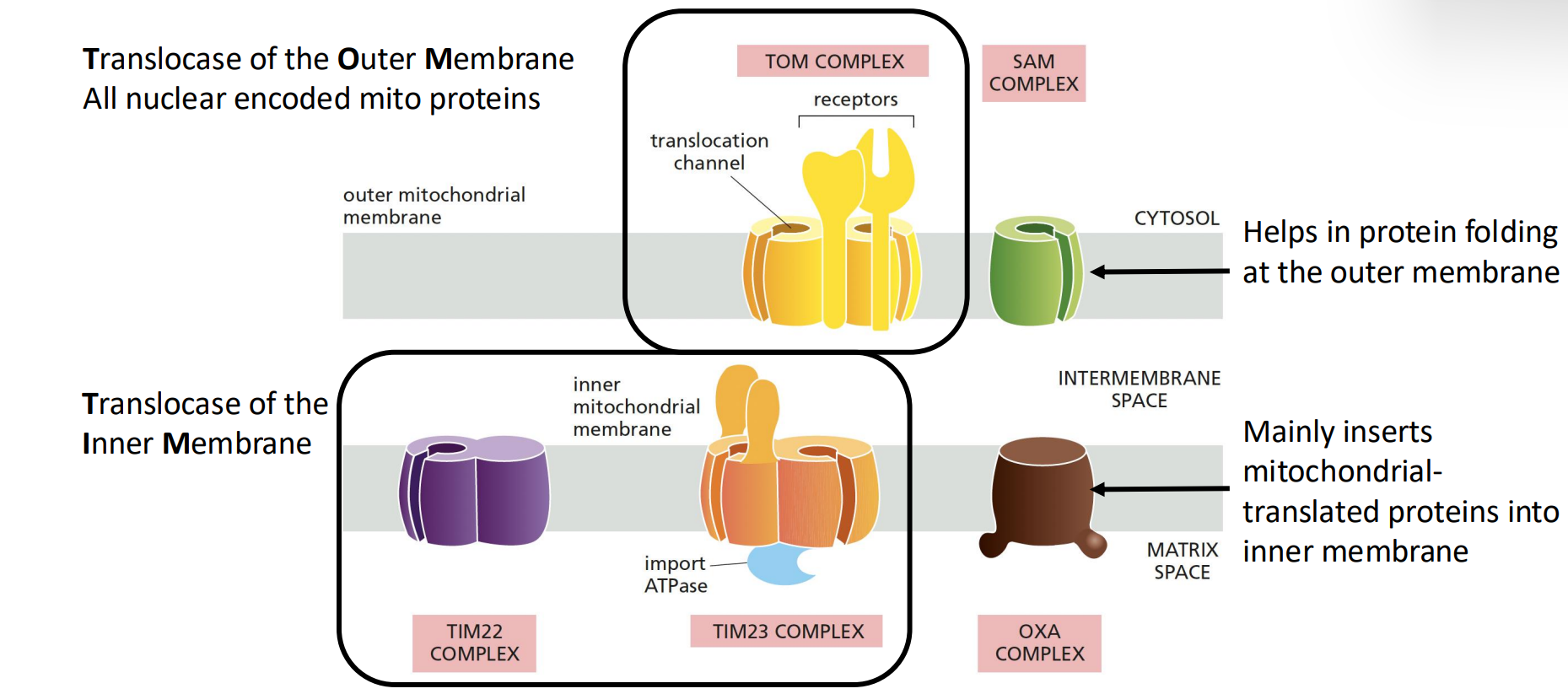
Translocase of the Outer Membrane (TOM complex):
Aids in protein folding at the outer membrane, preventing aggregation and misfolding.
SAM Complex: helps in protein folding at the outer membrane
Handles all nuclear-encoded mitochondrial proteins, ensuring that they are properly targeted and imported.
Translocase of the Inner Membrane (TIM complex):
TIM22 does transport of proteins, and TIM23 does mostly ATP
Primarily inserts mitochondrial-translated proteins into the inner membrane, facilitating their integration and function.
Oxa complex: involved in the insertion of proteins that are synthesized within mitochondria directly into the inner mitochondrial membrane.
Process of Soluble Protein Import into Mitochondria
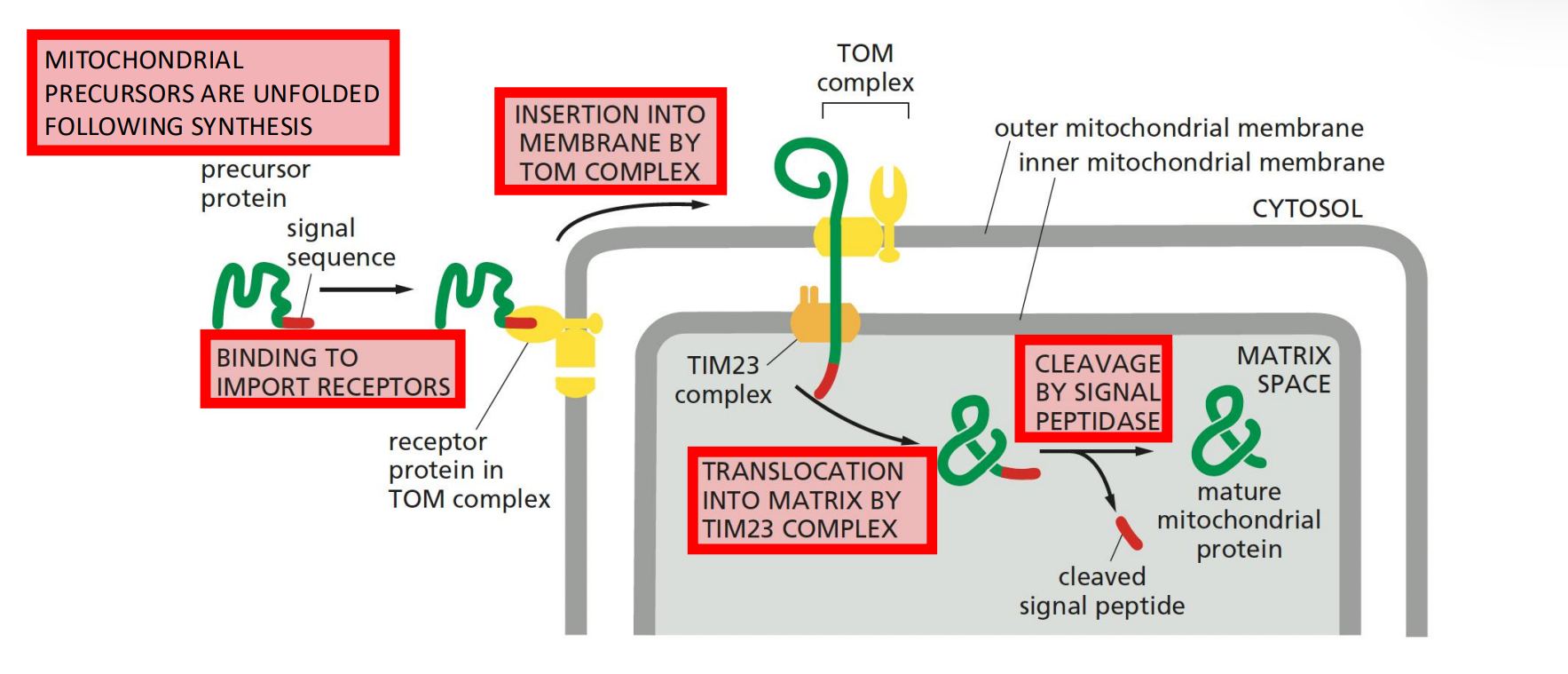
Mitochondrial precursors are unfolded following synthesis, allowing them to pass through the translocators. This is done through the chaperons hsp70
It is important to unfold the proteins because they won’t fit, but there also have to be factors that take care of the hydrophobic regions of the precursor proteins to make sure that they are not exposed to the outside of the protein interior
Binding to import receptors on the TOM complex initiates the import process.
Insertion into membrane by TOM complex facilitates the transfer of the protein across the outer membrane.
Translocation into matrix by TIM23 complex & cleavage by signal peptidase removes the MTS, allowing the protein to fold and function within the matrix.
The cleavage is important because therefore it cannot be exported from the matrix.
Energy Requirements for Mitochondrial Import
ATP hydrolysis and a membrane potential drive protein import to the mitochondria, providing the necessary energy for translocation and folding.
ATP hydrolysis by chaperones helps to a) disconnect the chaperons and unfold and b) pull proteins through the translocators.
Inner membrane potential (electrochemical gradient) assists in the translocation of positively charged MTS across the inner membrane.
mHsp60 assists in protein folding within the matrix, ensuring proper conformation and function.
Transport of Non-soluable Membrane Proteins into the Inner Mitochondrial Membrane
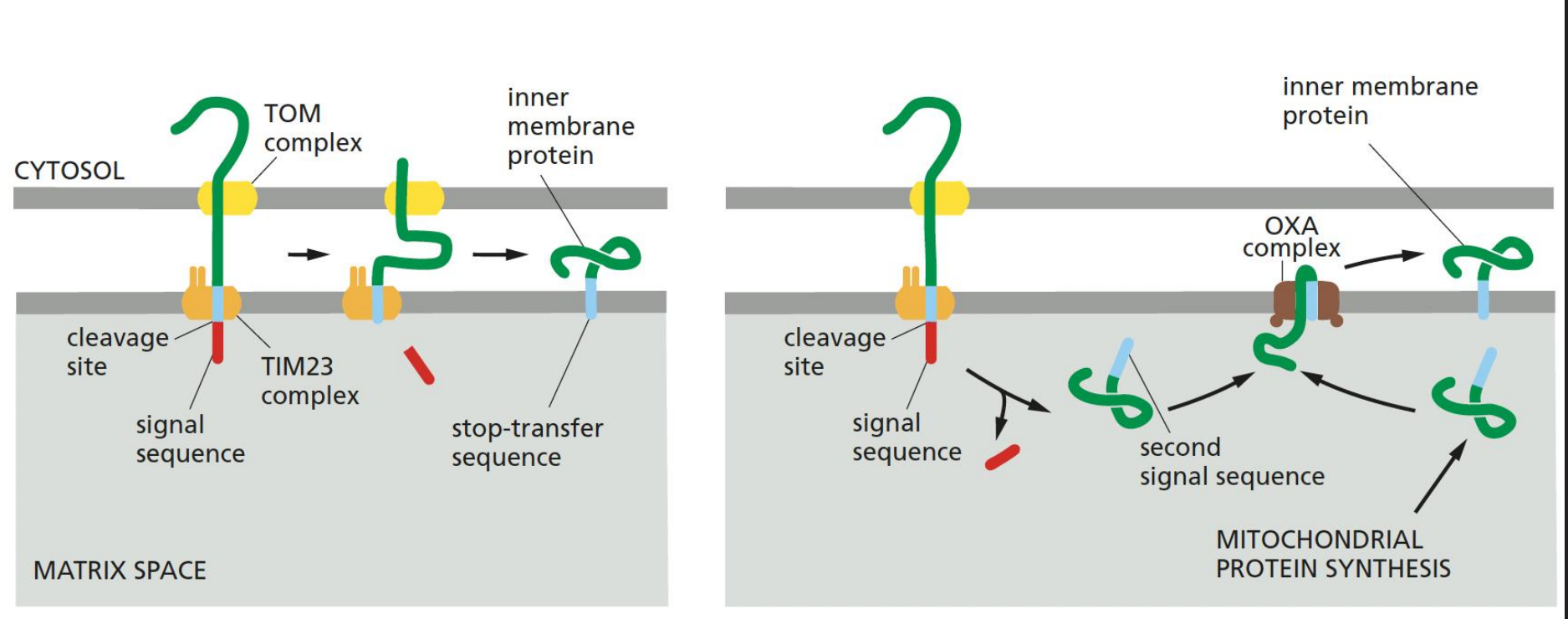
Requires a 'stop-transfer sequence' that halts translocation and anchors the protein in the membrane.
OXA complex is involved in mitochondrial protein synthesis and insertion of inner membrane proteins.
Second Signal Peptidase (release of Soluble Proteins into the Innermembrane space)
A second signal peptidase in the intermembrane space can remove the anchored hydrophobic sequence from soluble proteins, allowing them to be released into the IMS.
Targeting and Sorting Signals (review)
Mitochondrial proteins' targeting and sorting signals can be cleaved, permanent, or located at the N-terminus, depending on the specific protein and its destination.
Destinations, translocators, targeting/sorting signals:
Matrix: TOM complex, TIM23 complex, N-terminal matrix-targeting signal
IM (Inner Membrane): TOM complex, TIM23 complex, N-terminal transmembrane domain
IMS (Intermembrane Space): TOM complex, N-terminal sorting signal, TIM23 complex, C-terminal sorting signal
Parkinson’s Disease and Mitochondrial Dysfunction
Parkinson’s Disease involves:
Degeneration of the sustantia niagra of the brain
Motor impairment, including tremors, rigidity, and bradykinesia.
Cognitive and psychiatric symptoms, such as depression, anxiety, and dementia.
Autonomic dysfunction, affecting functions like blood pressure, digestion, and bladder control.
Linked to mitochondrial dysfunction and α-synuclein aggregation in dopaminergic neurons, contributing to the pathogenesis of the disease.
Characterized by selective loss of dopamine neurons in the substantia nigra and Lewy body formation, leading to motor deficits and other symptoms.
Alpha-Synuclein and Lipid Binding
Upon lipid binding, α-synuclein undergoes a conformational change, shifting to a highly α-helical conformation (71% α-helix when bound to lipid), which may be important for its function at the synapse.
Question: Does α-synuclein interact with TOM20? Because of the similarity of the alpha helix to a protein associated with it. This could provide insights into the mechanisms of synuclein aggregation and mitochondrial dysfunction in Parkinson's disease.
Observation: Positively charged amino acids clustered on one face of the α helix and nonpolar amino acids on the opposite side of import receptor, which may facilitate interaction with lipid membranes and other proteins.
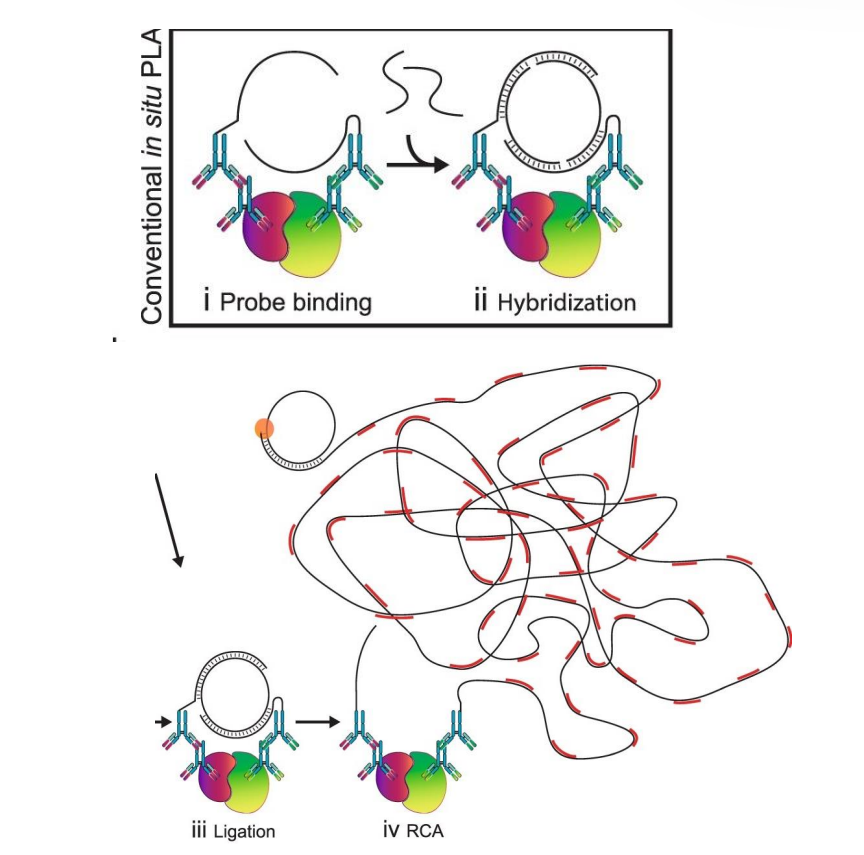
Proximity Ligation Assay (PLA) To Detect Protein Interaction
Antibodies are conjugated to oligonucleotides sequences. This enables specific binding to the target proteins and subsequent detection of their proximity.
In the proximity of less than , the addition of two oligonucleotides connector hybridize with the probes and form a closed circle template in the presence of a ligase. This indicates that the target proteins are in close proximity.
Addition of a polymerase and fluorescent-labelled oligonucleotides results in a rolling circle amplification of the closed circle template. This amplifies the signal, allowing for enhanced visualization of the protein interactions in cellular environments, which can provide critical insights into their functional dynamics.