Biochemistry term 4 week 2
Amino Acids
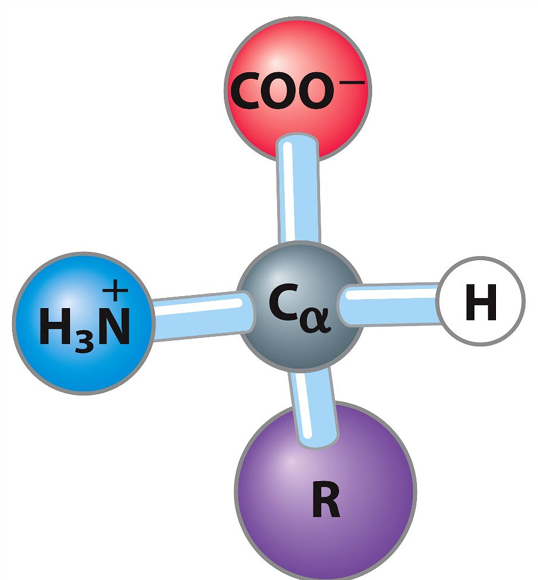
α-Amino Acids: Amino acids where the amino group is attached to the α-carbon atom.
Chirality:
Most α-amino acids are chiral, meaning they have a non-superimposable mirror image (enantiomer) due to the tetrahedral α-carbon bonded to four different groups.
Zwitterions:
Amino acids exist as zwitterions in solution, which are dipolar ions with both positive and negative charges.
pH-Dependent Charge:
The charge of an amino acid varies with pH due to the protonation and deprotonation of the amino and carboxyl groups.
pKa and pKR:
pKa is the dissociation constant for each ionizable group in the amino acid.
pKR is the dissociation constant for the R-group (side chain), if ionizable.
Isoelectric Point (pI):
The pH at which the amino acid has no net charge.
Calculation: where and are the pKa values of the carboxyl and amino groups, respectively (for amino acids without ionizable side chains).
Amino Acid Classification:
Amino acids are grouped based on the properties of their R-groups (e.g., nonpolar, polar, charged).
Amino Acid Structures and Properties
Nonpolar, Aliphatic R Groups:
Glycine, Alanine, Proline, Valine, Leucine, Isoleucine, Methionine.
Aromatic R Groups:
Phenylalanine, Tyrosine, Tryptophan
Note: Histidine is present in some slides under aromatic classification despite also being a positively charged R group.
Polar, Uncharged R Groups:
Serine, Threonine, Cysteine, Asparagine, Glutamine.
Positively Charged R Groups:
Lysine, Arginine, Histidine.
Negatively Charged R Groups:
Aspartate, Glutamate.
Titration Curves and Amino Acid Properties
Titration Curve Assignment Explanation
The titration curve of an amino acid can be analyzed to determine its pKa values and isoelectric point (pI).
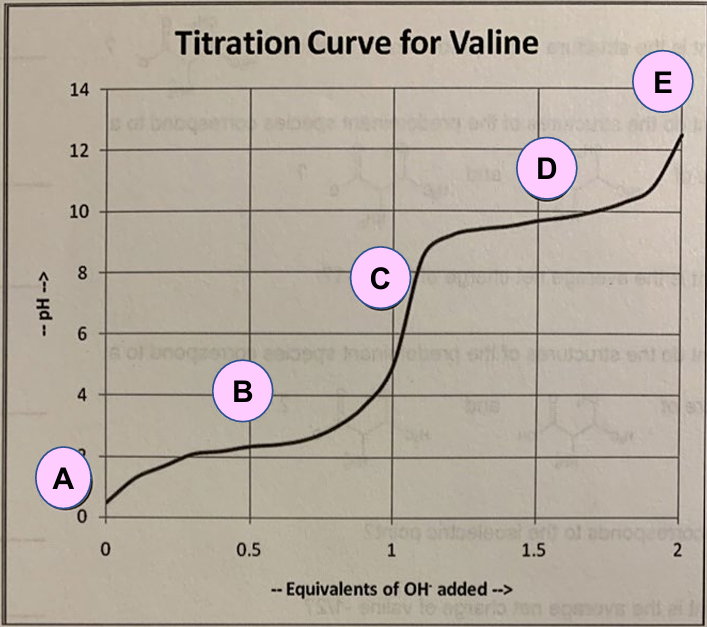
For Valine:
A: Predominant form of valine at low pH.
C: Predominant form of valine at high pH.
E: Predominant form of valine at a very high pH.
B: Indication of the first pKa (pK1).
D: Indication of the second pKa (pK2).
C: Indication of the pI.
pKa Values:
: pK of the carboxyl group
: pK of the amino group
Isoelectric Point (pI) Calculation:
For Valine:
Titration Curves of Arginine and Aspartic Acid
Arginine Titration Curve:
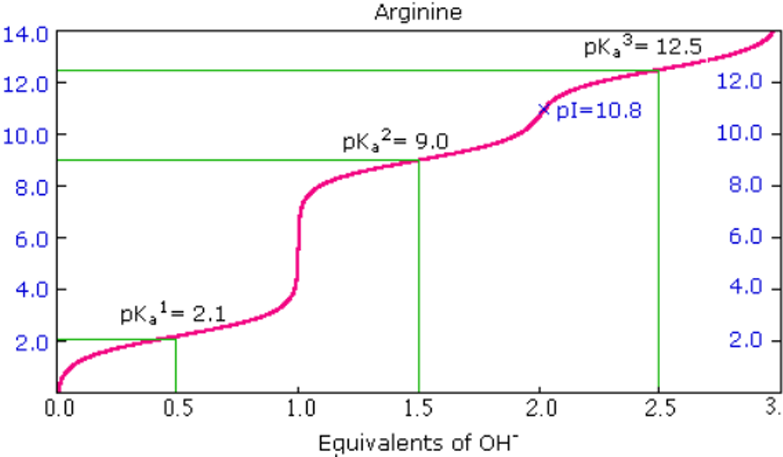
pKa1 = 2.1
pKa2 = 9.0
pKR3 = 12.5
pI = 10.8
Aspartic Acid Titration Curve:
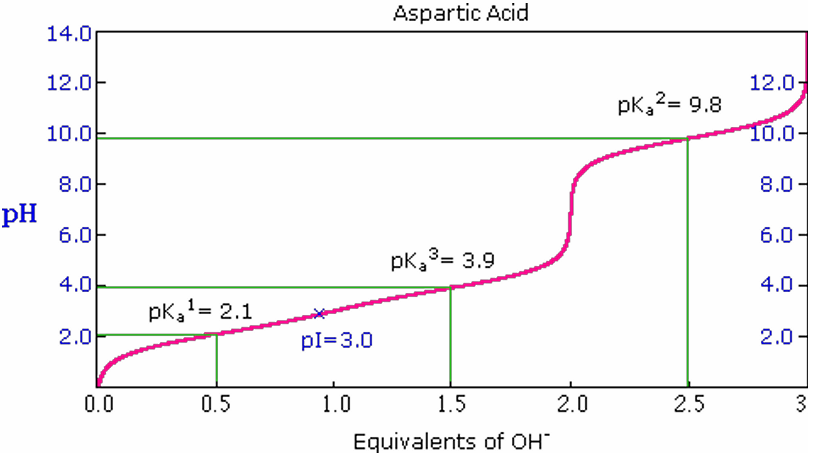
pKa1 = 2.1
pKa2 = 9.8
pKR3 = 3.9
pI = 3.0
Acidic vs. Basic Side Chains:
Negatively charged R groups (acidic side chains) are negatively charged if the pH is above the pKR value.
Positively charged R groups (basic side chains) are positively charged if the pH is below the pKR value.
Four Levels of Protein Structure
Primary Structure: Refers to the sequence of amino acids in a polypeptide chain.
Secondary Structure: Local spatial arrangement of the polypeptide chain's backbone atoms, commonly including α-helices and β-sheets.
Tertiary Structure: The overall three-dimensional structure of a polypeptide, resulting from interactions between R groups of amino acids.
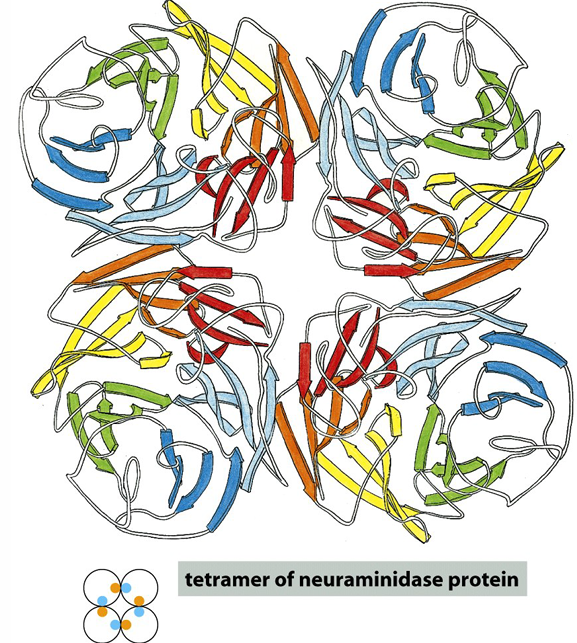
Quaternary Structure: The spatial arrangement of multiple polypeptide subunits in a protein complex.
Primary Structure: Peptides
Peptides: Small condensation products of amino acids formed through condensation reactions.
Naming Peptides:
Start at the N-terminus.
Full Amino Acid Names: serylglycyltyrosylalanylleucine
Three-Letter Code Abbreviation: Ser-Gly-Tyr-Ala-Leu
One-Letter Code (Proteins): SGYAL
Polypeptides: Charge and Isoelectric Point
Charge Prediction: Based on the pK of each group involved, the charge at a defined pH can be predicted.
Isoelectric Point (pI) Determination: The algorithm to determine the pI of a polypeptide chain can be found in databases. => pI= (pKa1+pKa2)/2
Exercise: Net Charge Calculation of a Peptide
Peptide Sequence: Gly-His-Trp-Glu-Gly-Leu-Arg-Pro-Gly
Assume pKa for the α-carboxyl group is 2 and for the α-amino group is 9.pKR Values of Charged Amino Acids:
Lysine (Lys, K): 10.5
Histidine (His, H): 6.0
Arginine (Arg, R): 12.5
Aspartate (Asp, D): 3.7
Glutamate (Glu, E): 4.3
Calculations:
pH 3:
H3N+, HN+, H2N+, COO-
Net charge: +2
pH 8:
H3N+, N, H2N+, COO-
Net charge: 0
pH 11:
H2N, N, H2N+, COO-
Net charge: -1
Primary Structure: Local Bending of the Chain
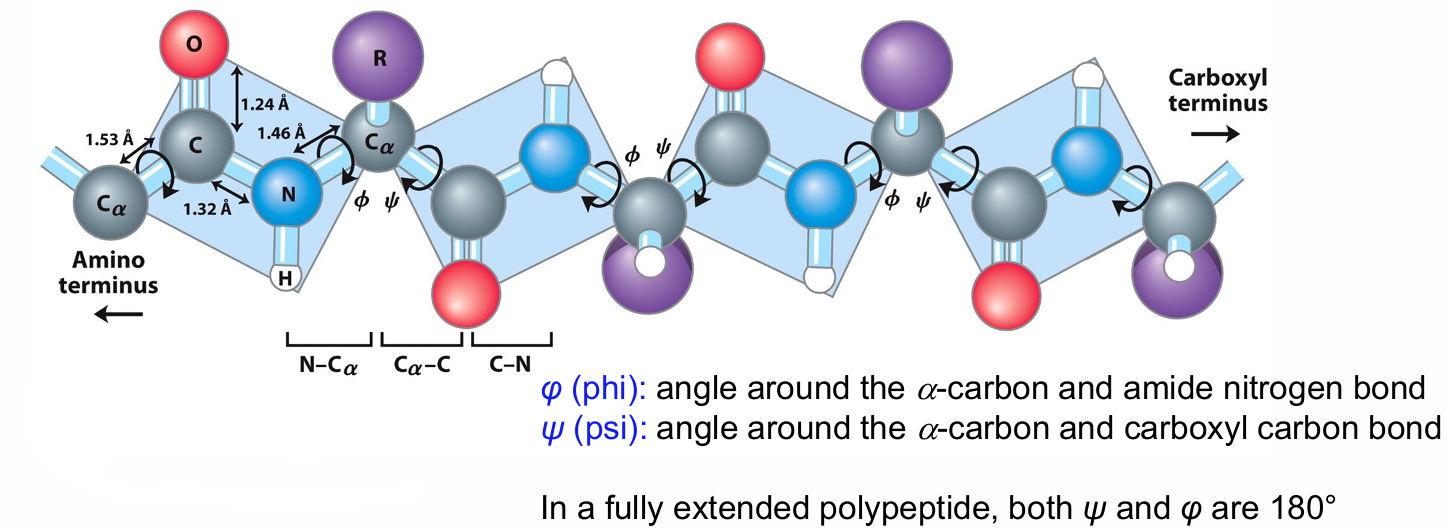
Peptide Bond Planarity: The six atoms around the peptide bond are in a single plane.
α-Carbons as Hinges: The Cα atoms serve as hinges between these planes.
Torsion Angles:
φ (phi): Angle around the α-carbon and amide nitrogen bond.
ψ (psi): Angle around the α-carbon and carboxyl carbon bond.
Fully Extended Polypeptide:
In a fully extended polypeptide, both ψ and φ are 180°.
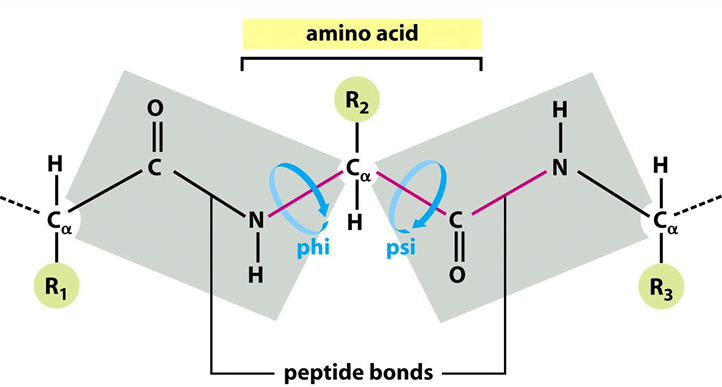
Steric Hindrance: Some φ and ψ combinations are unfavorable due to steric hindrance.
Favorable H-bonding: Some φ and ψ combinations are more favorable due to the chance to form favorable H-bonding interactions along the backbone.
Secondary Structures
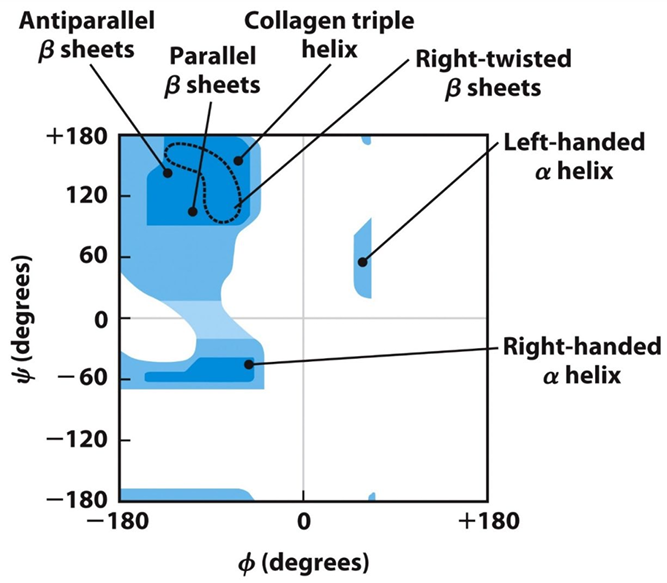
Ramachandran Plot: Shows the distribution of φ and ψ dihedral angles found in a protein.
Definition: Local spatial arrangement of main-chain atoms (backbone of the polypeptide).
Common Regular Arrangements:
α-Helix: Stabilized by hydrogen bonds between the peptide backbone of nearby amino acids.
β-Sheet: Stabilized by hydrogen bonds between peptide backbone spatially adjacent segments that may not be nearby in the peptide chain.
Random Coil: Irregular arrangement of the polypeptide chain.
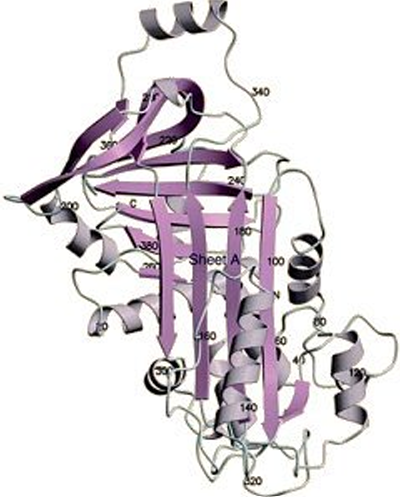
Secondary Structure: α-Helix
Structure: Tightly coiled backbone forming the inner part, with side chains extending outward.
Stabilization: Stabilized by hydrogen bonds between the CO group and the NH of the fourth residue ahead.
Destabilizing Amino Acids:
Long stretches of Glu, Arg, or Lys
Val, Thr, Ile, Ser, Asp, Asn
Pro, Gly
Stabilizing Amino Acids:
Ala, Leu
Common Helix Type:
The right-handed helix is the common α-helix (3.6 residues/turn).
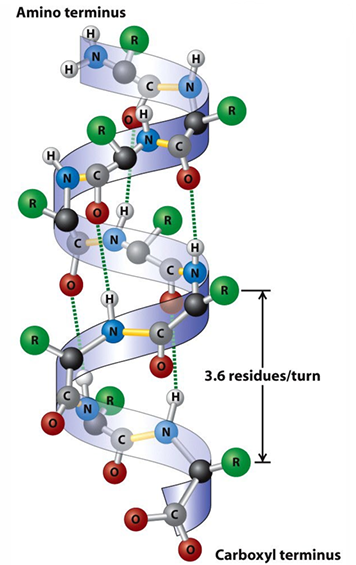
Secondary Structure: β-Sheet
Pleated Sheet: The backbone polypeptide chain is extended into a zig-zag, with side chains protruding from the strand alternating in up and down directions.
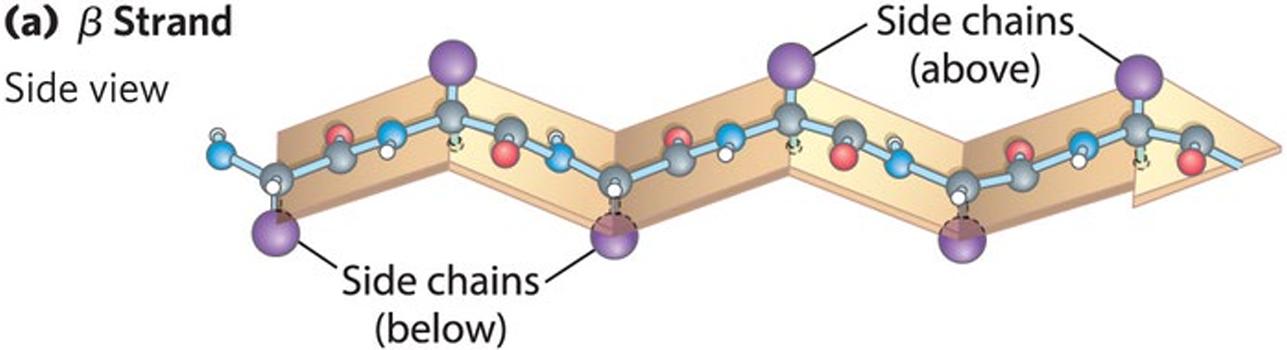
Arrangement: Held together by hydrogen bonds between backbone amides in different strands.
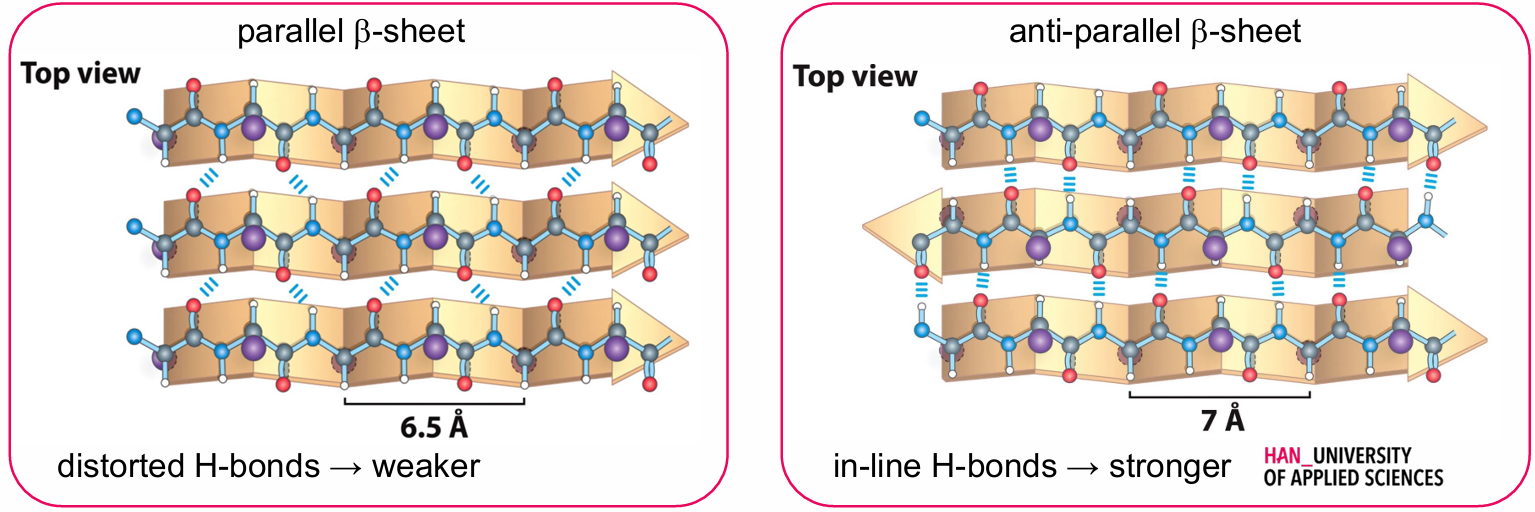
Types:
Parallel β-Sheet
Anti-Parallel β-Sheet
Hydrogen Bonds:
Distorted H-bonds → weaker
In-line H-bonds → stronger
Secondary Structure: Loops
Function: Connect α-helices and β-sheets, usually causing a change in direction.
Properties: Hydrophilic, on the surface of proteins, and not well-defined.
Turns: Loops consisting of only 4-5 amino acids forming internal hydrogen bonds.
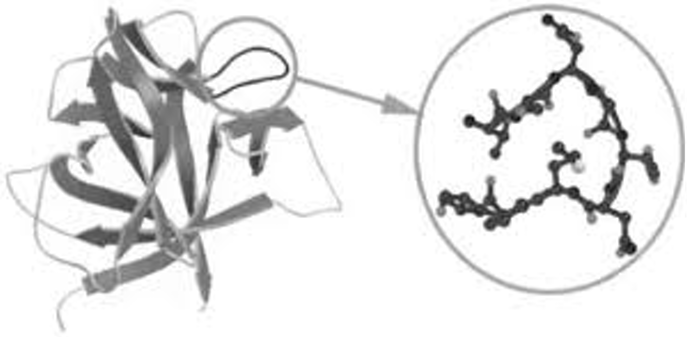
Secondary Structure: β-Turns
Frequency: Occur frequently whenever strands in β sheets change direction.
Structure: The 180° turn is accomplished over four amino acids.
Stabilization: Stabilized by a hydrogen bond from a carbonyl oxygen to an amide proton three residues down the sequence.
Common Amino Acids:
Proline in position 2 or Glycine in position 3 are common in β turns.
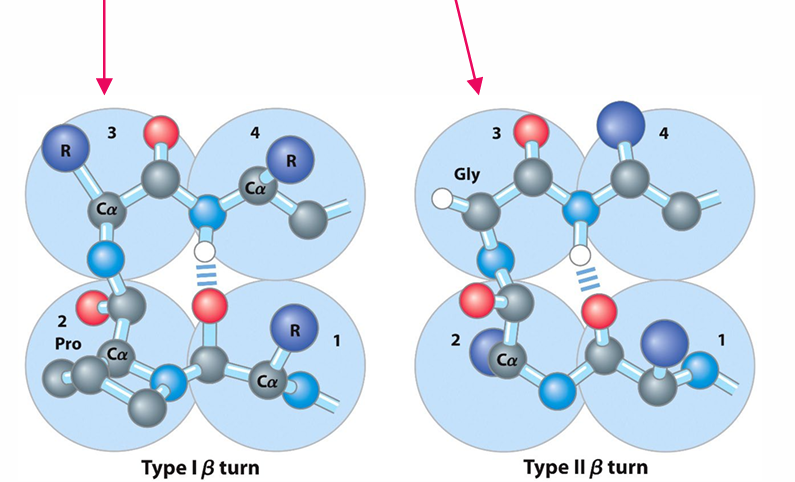
Peptide Bonds: Most peptide bonds many proline-peptide bonds in β-turns
Tertiary Structure
Definition: The overall 3D structure of a polypeptide.
Formation: Primarily due to interactions between the R groups of the amino acids that make up the protein.
Note: Interacting amino acids are not necessarily next to each other in the primary sequence.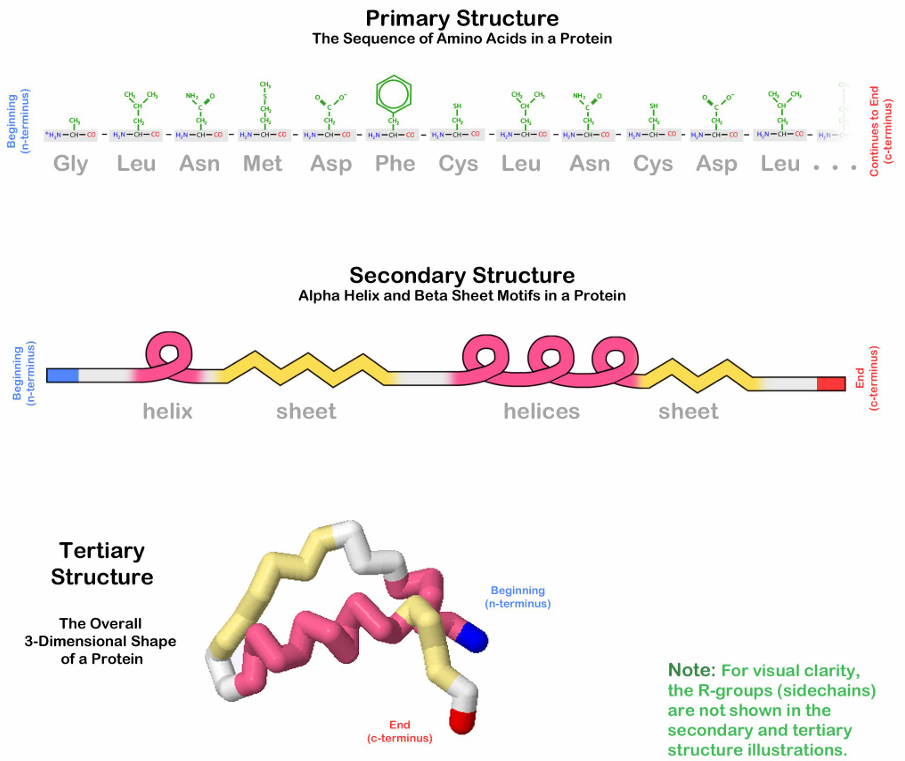
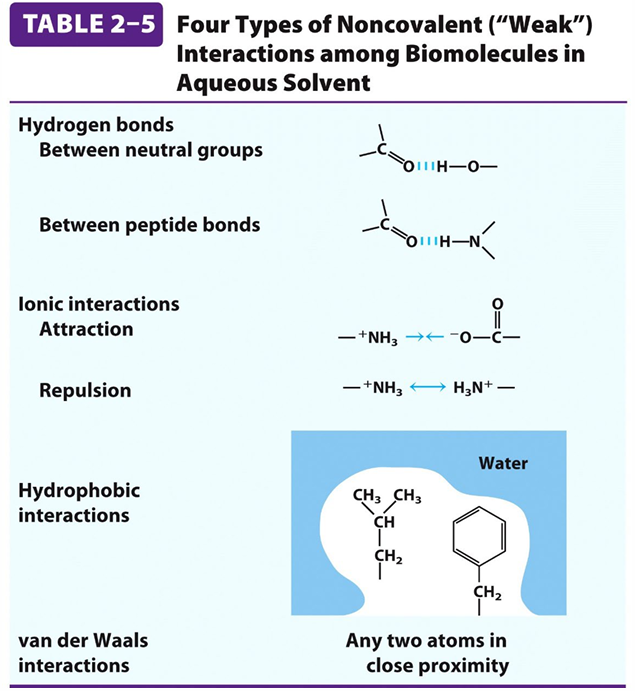
Interactions: Formed by various noncovalent interactions and disulfide bonds.
Ionic bond
Hydrogen bond
Disulfide linkage
Hydrophobic interactions
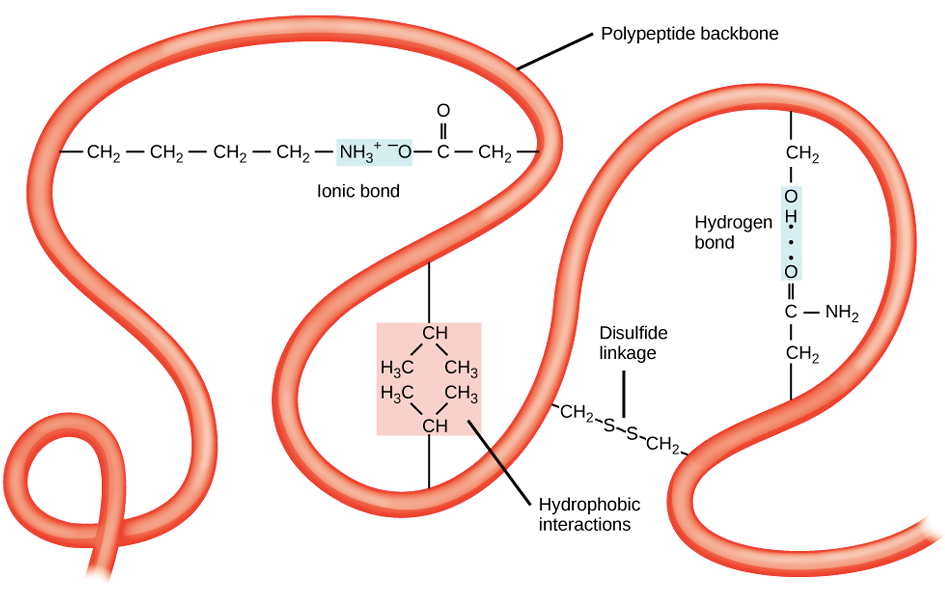
Tertiary Structure: Examples of Noncovalent Interactions
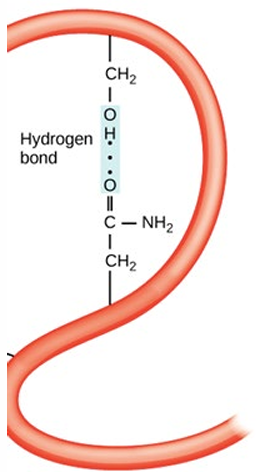
Polar Side-Chains: Interactions between polar side-chains.
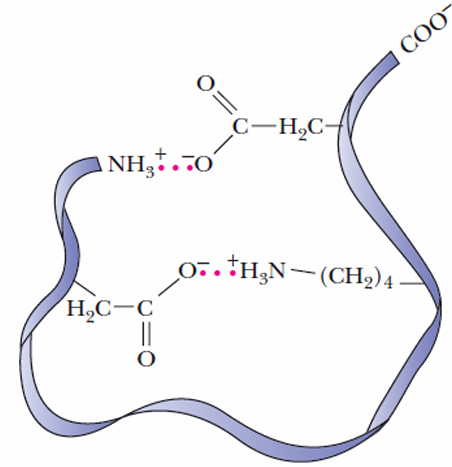
Charged Amino Acids (Electrostatic Interactions):
Types of Interactions:
Salt Bridges: Occur between oppositely charged amino acids like lysine and aspartate. These are strong interactions contributing significantly to protein stability.
Ion-Dipole Interactions: Involve interactions between an ion and a polar amino acid residue.
Dipole-Dipole Interactions: Occur between polar amino acids, aligning to maximize attractive forces.
Environmental Influence: The strength of electrostatic interactions can be affected by the surrounding environment, such as pH levels and the presence of other ions.
Hydrophobic Side-Chains Interactions:
Mechanism: Hydrophobic amino acids such as valine, leucine, and isoleucine tend to cluster together in the protein's interior to avoid contact with water. This hydrophobic effect is a primary driving force in protein folding.
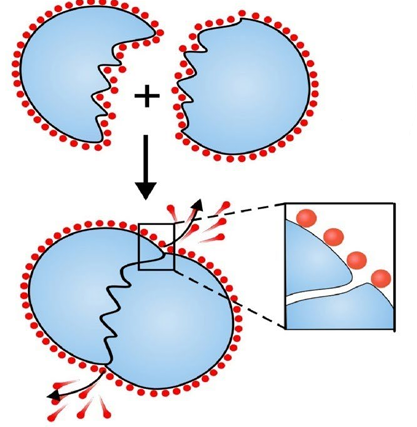
Stability: These interactions are stabilized by van der Waals forces between the hydrophobic residues, contributing to the overall stability of the protein.
Weak Attractions (Van der Waals Forces):
Description: These are medium-range, weak attractions that occur between all atoms. They include London dispersion forces, dipole-dipole interactions, and dipole-induced dipole interactions.
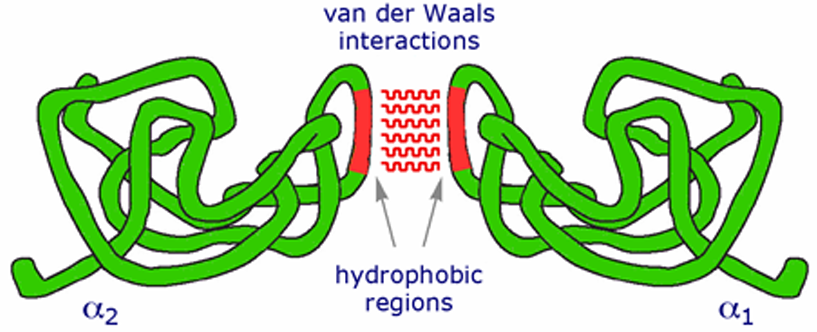
Significance: Although each interaction is weak, the cumulative effect of numerous van der Waals forces significantly contributes to the stability of the protein's interior. This is crucial for maintaining the protein's overall three-dimensional structure, ultimately influencing its function.
Tertiary Structure: Disulfide Bonds
Formation: Covalent bond between R groups of cysteine residues.
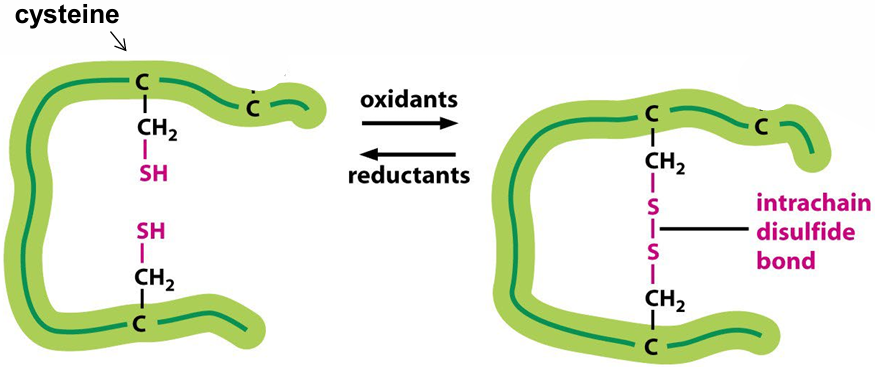
Quaternary Structure
Definition: Proteins with more than one polypeptide chain exhibit a fourth level of structural organization.
Arrangement: The spatial arrangement of subunits (chains) and the nature of their interactions based on interactions between rest groups (as for tertiary).
Interactions:
Hydrogen bonds
Electrostatic interactions
Van der Waals interactions
Hydrophobic interactions
Disulfide bonds