Translation and Protein Synthesis
Translation Overview
Site: Translation occurs in the cytosol, cytoplasm.
Process: mRNA base-pair sequence is translated to amino acid (aa) sequence.
Codon Reading: Nucleotides read in groups of three, known as codons.
Reading Frames: Each reading frame can lead to a different amino acid sequence, emphasizing the importance of the starting point in translation. In principle, an RNA sequence can be translated in any one of three different reading frames.
Function: Set to Start; Set to Stop ✋
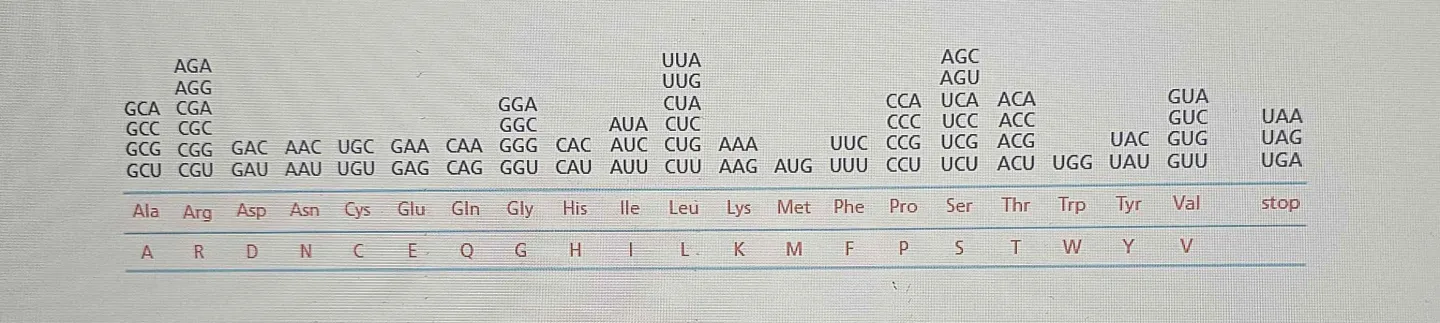
Genetic Code: The relationship between codon and its corresponding amino acid.
Codons: 64 codons (4 different bases x 3 bases per code = 64) , which specify 20 different amino acids, along with start and stop signals that regulate the translation process. The start codon, typically AUG, signals the beginning of translation, while stop codons (UAA, UAG, UGA) indicate the termination of the polypeptide chain.
RNA is a linear polymer of four different nucleotides, so there are
4 × 4 × 4 = 64 possible combinations of three nucleotides: the triplets AAA, AUA, and AUG. These combinations correspond to the 20 standard amino acids protein…
1. Some triplets serve as codons for the same amino acid or are specified by more than one triplet., which leads to redundancy in the genetic code. This redundancy allows for mutations to occur without necessarily altering the resulting protein, providing stability to the genetic information.
2. Either some nucleotide triplets are never used, or the code is redundant
3. The correct one: Each group of three consecutive nucleotides in RNA is called a codon, and each codon specifies either one amino acid or a stop to the translation process.
For example:
GCU, GCC, GCA, GCG → All code for Alanine (Ala).
AAA, AAG → Both code for Lysine (Lys).
Even though there are duplicate (synonymous) codons, each one is locked into coding only that one amino acid. No codon ever flips to mean something else.
One specific amino acid. This precise relationship between codons and amino acids is fundamental to the genetic code, allowing for the accurate synthesis of proteins during translation.️
Base pairing
There is more than one tRNA for many of the amino acids, and some tRNA molecules can base-pair with more than one codon or both.
What is Wobble Base Pairing?
The standard base-pairing rules apply strictly for the first two nucleotides of a codon.
The third position is more flexible, allowing some non-standard pairings (hence, the "wobble").
This flexibility reduces the number of tRNAs needed to read all 61 sense codons. The remaining 3 codons (UAA, UAG, UGA) are stop codons, which signal the end of translation.
As a result, most organisms have far fewer than 61 tRNAs—usually around 40-45 tRNAs are enough to decode all the sense codons.
Why Does Wobble Matter?
Redundancy of the Genetic Code – Many amino acids are specified by multiple codons that differ only in the third base.
Efficiency in Translation – Cells don’t need 61 different tRNAs; instead, some tRNAs can recognize multiple codons.
When a tRNA "scans" mRNA, it can recognize different codons that code for the same amino acid—it doesn't need to worry about which specific codon it encounters.
So, regardless of the variation in the third base (wobble), the tRNA knows it means the same thing: the same amino acid.
Mutational Buffering – Mutations in the third codon position often don’t change the amino acid (silent mutations), reducing harmful effects.
EX: tRNA base (Anticodon) → mRNA base (Codon)
G → U or C
U → A or G
I (Inosine) → U, C, or A
This explains why the genetic code is degenerate but not ambiguous—multiple codons can specify the same amino acid, but each codon still unambiguously codes for only one amino acid! 🚀
Degenerate (Redundant): Multiple codons can specify the same amino acid.
Example: GCU, GCC, GCA, and GCG all code for Alanine (Ala).
Not Ambiguous: Each codon always codes for just one amino acid—it never switches meaning.
Example: GCU will always mean Alanine, never something else.
This setup is efficient and protective:
✅ Redundancy helps prevent harmful effects from mutations.
✅ Unambiguity ensures accuracy in protein production.
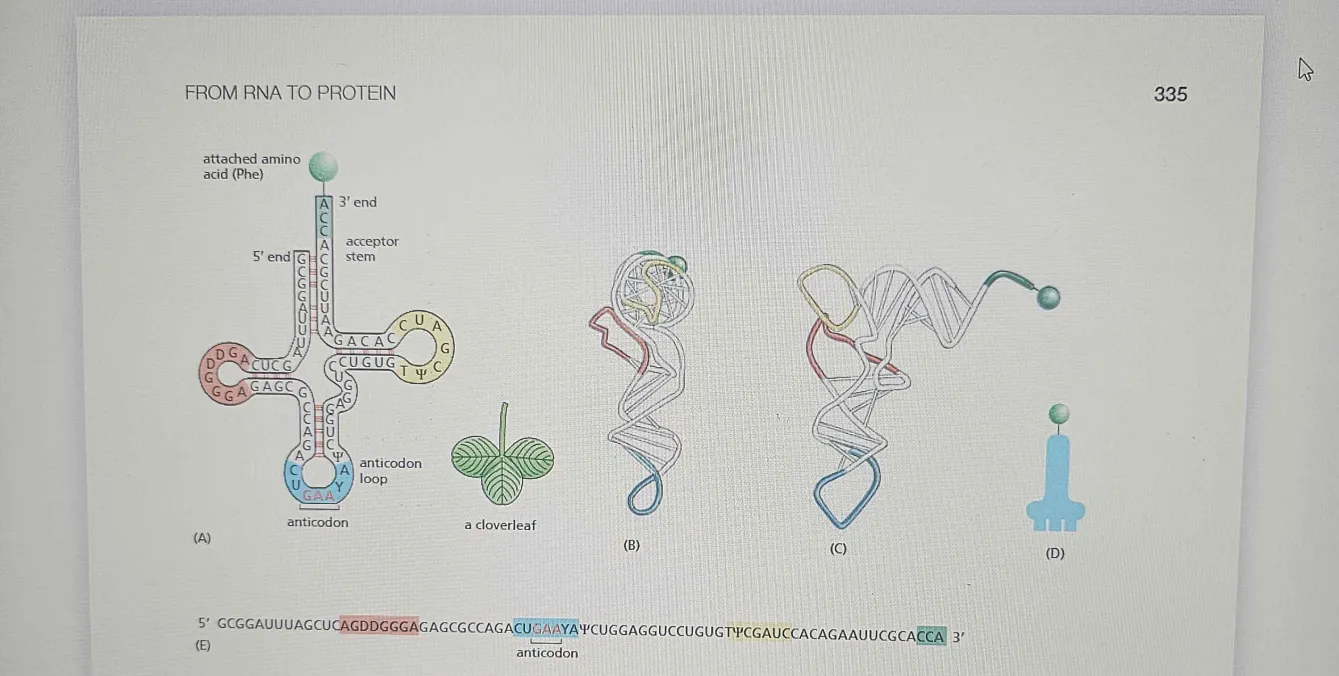
tRNA Molecule
tRNA molecules act as adaptors by recognizing specific codons in the mRNA sequence and delivering the corresponding amino acids during protein synthesis. Each tRNA has two critical regions:
Function:
Links codon on mRNA to amino acid.
Anticodon: A set of three nucleotides that are complementary to a specific mRNA codon. This ensures the correct amino acid is added to the growing polypeptide chain.
Amino Acid Attachment Site: At the 3' end of the tRNA, where a specific amino acid is covalently attached by an enzyme called aminoacyl-tRNA synthetase (which ensures the correct pairing).
Size: Average of 80 nucleotides.
Structure: Contains an anticodon region that binds to the codon on mRNA.
tRNAs themselves are not amino acids, but they mimic the shape and function of the amino acid they carry.
tRNA Characteristics
Shape:
four short segments of the folded tRNA are double-helical, producing a molecule that looks like a cloverleaf when drawn schematically
tRNA folds into a cloverleaf shape due to complementary base pairing between different molecule regions.
L-Shaped 3D Structure (Tertiary Structure) 🏗
The cloverleaf undergoes further hydrogen bonding and stacking interactions, forming a compact L-shape in 3D.
This structure is crucial for fitting into the ribosome during translation.
One of these regions forms the Anticodon, a set of three consecutive nucleotides that pairs with the complementary codon in an mRNA molecule. The other is a short single-stranded region at the 3ʹ end of the molecule; this is the site where the amino acid that matches the codon is attached to the tRNA
Trna+Trna=Stem
Base-pairing between complementary regions of the same tRNA forms the (double-stranded).
The stem in tRNA is made up of base-pairing between complementary regions of the tRNA itself (i.e., between two complementary strands of the same tRNA), not between mRNA codons
(like A-U and G-C).
mRna Codon + Anticodon = Loop: The loops are where you get single-stranded regions. The remaining unpaired regions (including the anticodon loop) form the loops.
If the mRNA codon is AUG, the tRNA’s anticodon will be UAC, and they’ll pair in the anticodon loop.
Transcription & Initial Processing
tRNAs Are Covalently Modified Before They Exit from the Nucleus
Eukaryotic tRNAs are transcribed by RNA polymerase III; they are long.
Bacterial and eukaryotic tRNAs are synthesized as precursor tRNAs, which must be trimmed to produce mature tRNAs.
tRNA Splicing (Differs from pre-mRNA Splicing!)
Processed by trimming at the ends and in the middle.
Some tRNA precursors contain introns that need to be spliced out.
Instead of a lariat intermediate (like mRNA splicing), tRNA splicing uses a cut-and-paste mechanism catalyzed by proteins.
Proper cloverleaf folding is required for trimming and splicing—misfolded tRNAs won’t be processed correctly, acting as a quality-control step.
Chemical Modifications (Essential for Function!)
Nearly 1 in 10 nucleotides in mature tRNA is a modified version of A, U, C, or G.
Over 50 different types of tRNA modifications exist!
Key Modification: Inosine (I)
Formed by deamination of adenosine (A).
Affects anticodon base-pairing, helping tRNA recognize multiple mRNA codons (wobble base-pairing).
Other modifications enhance stability, folding, and amino acid attachment accuracy. Synthesized by RNA polymerase III as large transcripts.
Chemical modifications produce unique non-traditional bases (Ψ, Im, Gm, etc.) that define a unique conformation.
It has three stem-loops (D loop, TψC loop, and anticodon loop) and an acceptor stem where the amino acid attaches.
Ψ (Pseudouridine): A modified form of uridine that enhances the stability and function of RNA.
Im (Inosine): Can be involved in wobble base pairing and is often found in tRNAs.
Gm (Methylguanosine): A methylated version of guanosine, often found in RNA caps.
I (Inosine): Found in tRNAs and is important for base-pairing flexibility.
T (Thymine): Occasionally used in RNA processing.
Why the tRNA is Trimmed:
Trimming at the Ends:
The 5' and 3' ends of the precursor tRNA have extra sequences that are not needed in the mature tRNA.
These extra nucleotides at the ends are removed to create the correct mature length and proper structure for the tRNA. This ensures that the tRNA will fold properly and be able to interact with other molecules like aminoacyl-tRNA synthetase and the ribosome.
Trimming in the Middle:
There is a portion of the precursor tRNA (the intervening sequence) that is also removed. This part is usually a spacer sequence that is not needed for the mature tRNA’s function.
The middle trimming removes this unnecessary sequence to allow the stem-loop structure to form properly, which is essential for the tRNA to function correctly during translation.
The Steps of tRNA Processing:
Transcription of Precursor tRNA:
RNA polymerase III transcribes a long precursor tRNA from the DNA template.
Trimming at the Ends:
Enzymes trim the 5' and 3' ends of the precursor tRNA to remove extra nucleotides that aren’t needed for the mature tRNA.
Splicing (Trimming in the Middle):
Some precursor tRNAs have an intervening sequence (a non-coding region) in the middle, which is spliced out (removed) by RNA splicing enzymes. This helps form the proper stem-loop structure.
Base Modifications:
After trimming, certain bases are chemically modified (e.g., pseudouridylation, methylation) to help the tRNA maintain its proper function and 3D shape.
Final Step:
The mature tRNA is now ready for translation, with a correct structure, including an anticodon loop and a 3' CCA tail for attaching the amino acid.
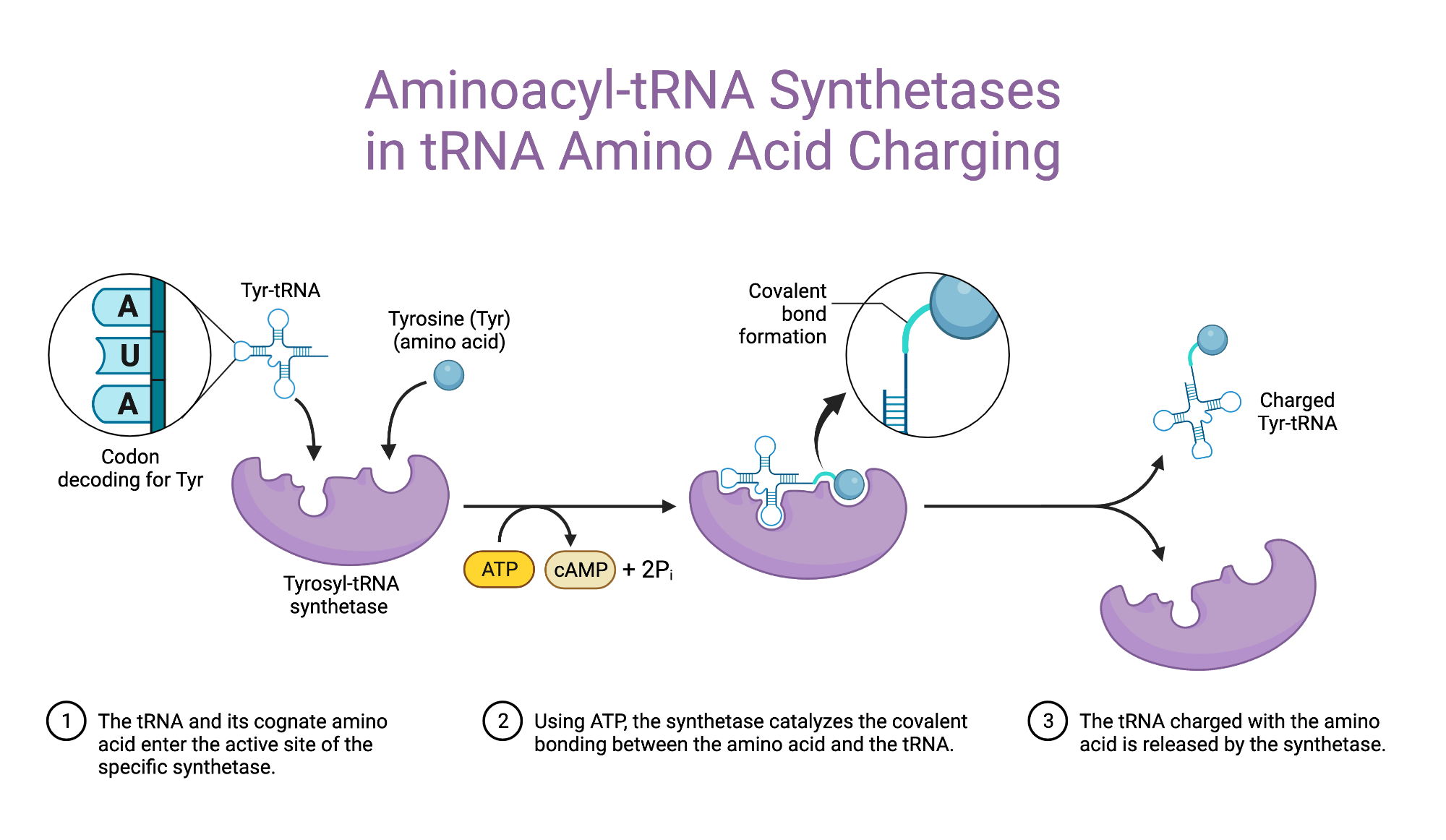
Aminoacyl Synthetase
Specific Enzymes Couple Each Amino Acid to Its Appropriate tRNA Molecule
Function: That has an anticodon.
Links amino acids to tRNA (e.g., tryptophan).
These enzymes covalently attach the correct amino acid to the corresponding tRNA.
This step is crucial for ensuring accurate translation of the genetic code!
Most cells have 20 different aminoacyl-tRNA synthetases, one for each of the 20 amino acids.
Each synthetase recognizes the correct amino acid and its matching tRNA(s).
Example: Glycyl-tRNA synthetase attaches glycine to all tRNAs that recognize glycine codons.
Some bacteria have fewer than 20 synthetases.
A single synthetase can sometimes attach the same amino acid to multiple tRNAs.One synthetase → Can charge multiple tRNAs if they correspond to the same amino acid.
One of these tRNAs may undergo further modification to ensure correct pairing during translation.
Error:
Normally, each aminoacyl-tRNA synthetase attaches the correct amino acid to its matching tRNA.
However, some bacteria have fewer than 20 synthetases, meaning one synthetase may attach the wrong amino acid to a tRNA.
Solution: A second enzyme chemically modifies the incorrect amino acid so that it now matches the anticodon of the tRNA.This ensures the correct amino acid is delivered to the ribosome during translation.
Energy:
The reaction attaching the amino acid to the 3′ end of the tRNA requires ATP (energy).
This reaction forms a high-energy bond between the amino acid and tRNA.
The reason the formation involves a high-energy bond, which is then utilized later to form peptide bonds, is because…
Later, during protein synthesis, this stored energy is used to covalently link the amino acid to the growing polypeptide chain in the ribosome.
Without this energy input, forming peptide bonds between amino acids would be much harder!
Reaction: Involves ATP breakdown to AMP + 2 Pi in the linkage process.
Aminoacyl-tRNA synthetases = Chefs who attach ingredients (amino acids) to the right containers (tRNAs).
Mistakes happen! If the wrong ingredient is placed in a container, a second helper corrects it (chemical modification).
If aminoacyl-tRNA synthetases make a mistake, the wrong amino acid gets placed into a protein, which can mess up its function—kind of like putting the wrong ingredient in a recipe 🍳.By having highly specific synthetases, cells reduce errors and ensure proteins are built correctly and efficiently!
ATP = Stove 🔥—providing the energy needed to cook (form peptide bonds).
High-energy bond = Pre-cut ingredients 🥕🔪—when it’s time to cook (protein synthesis), the hard part is already done, making the process fast and efficient!
The high-energy bond is formed when an amino acid is attached to tRNA using ATP energy (like pre-cutting veggies).
This stores energy in the bond between the amino acid and tRNA, like prepping ingredients ahead of time.
When it’s time for translation (protein synthesis), the ribosome doesn’t need extra energy—it can just use the already activated amino acid to build the protein quickly and efficiently.

Understanding Aminoacyl-tRNA Synthetases 🛠
Think of tRNAs as delivery trucks 🚚 that carry amino acids to the ribosome factory 🏭 to make proteins. But before these trucks can deliver anything, they need to be loaded with the correct cargo (amino acid).💡 Here’s where aminoacyl-tRNA synthetases come in—they act like warehouse managers 📦. Each manager is responsible for checking the truck (tRNA) and loading it with the right cargo (amino acid). Aminoacyl-tRNA synthetases are like quality control managers who make sure every tRNA (truck) gets loaded with the correct amino acid (cargo) before heading to the ribosome (factory). 🚛📦🏭
Molecular Proofreading
First discrimination step:
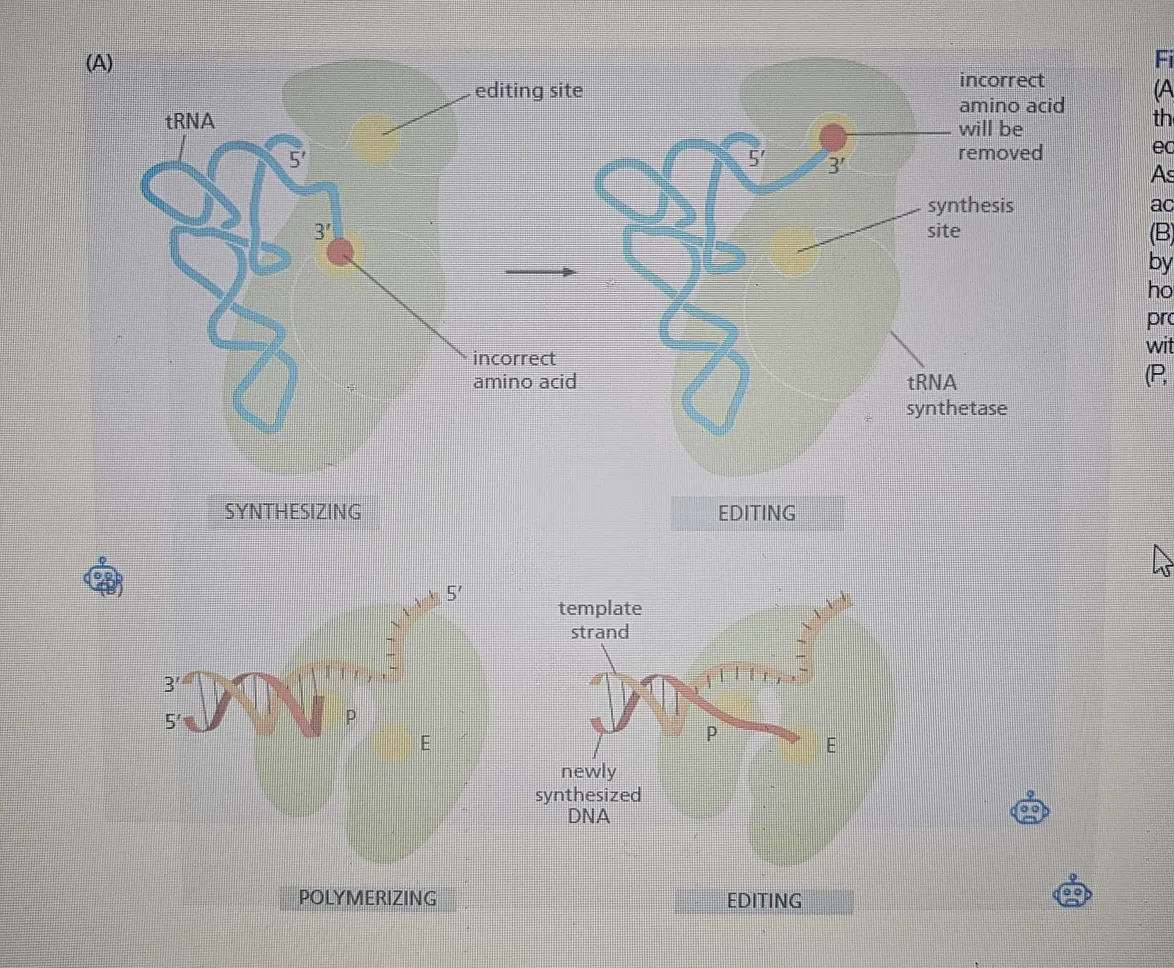
The synthetase uses its active-site pocket preferentially (HIGH AFFINITY for the site), thus binding the correct amino acid, favoring it over others (19) based on size and shape. The correct amino acid fits best, while larger amino acids are physically excluded from the pocket. This step helps reduce errors, but it’s not enough when amino acids are very similar, like valine and isoleucine ( which differ by only a methyl). Amino acids are difficult to achieve in a single step.
Second discrimination step (editing):
After the amino acid is covalently linked to AMP, the synthetase enters a second phase. It binds tRNA, and the adenylated amino acid is forced into an editing pocket within the enzyme. The editing pocket has very specific dimensions that prevent the correct amino acid from fitting, but it allows closely related amino acids like valine and isoleucine to access it. This allows the synthetase to “proofread” the amino acid by hydrolyzing (removing) the incorrectly attached amino acid before it gets linked to the tRNA.
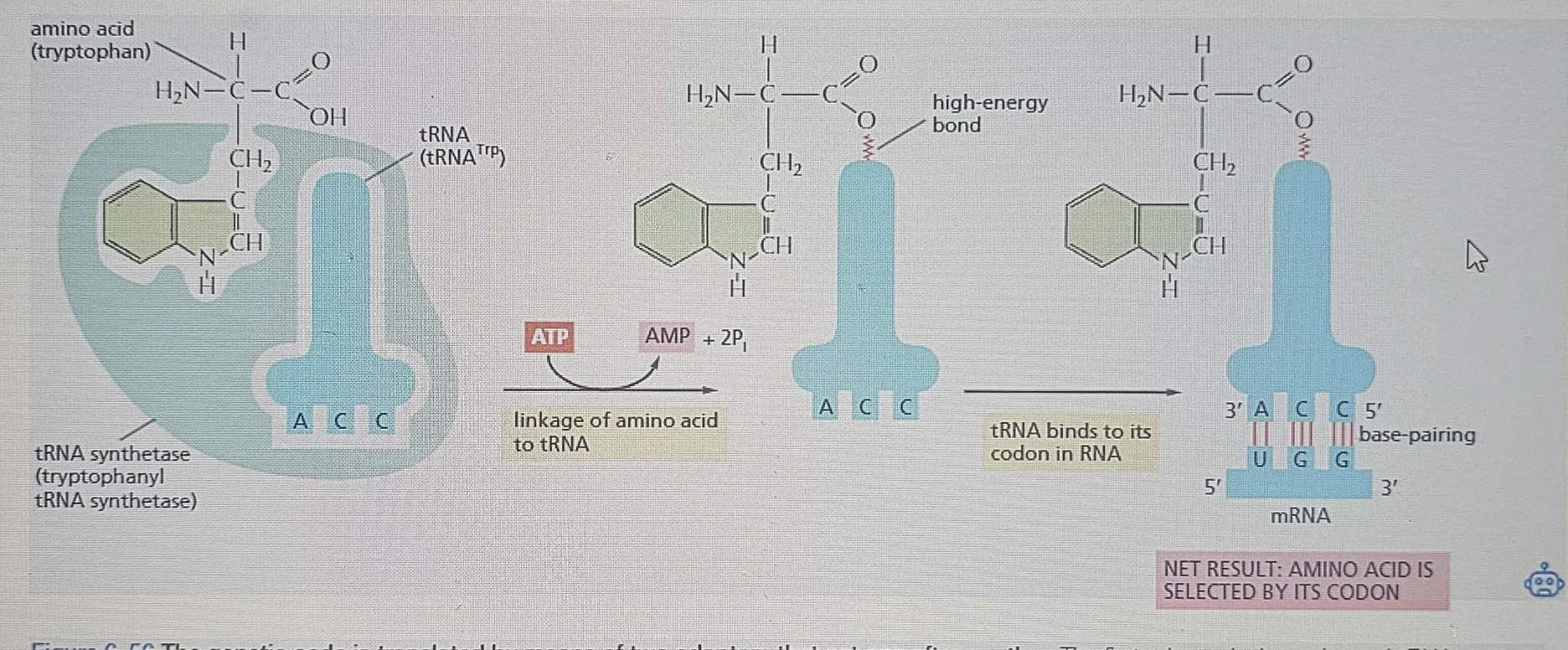
Hydrolytic editing:
This hydrolysis process is like exonucleolytic proofreading in DNA polymerases, where mistakes (such as a wrong nucleotide being added) are removed. The final product is an accurately charged tRNA, with errors reduced to about one mistake in 40,000 couplings, which is impressively accurate.
Activation outside the editing site:
The amino acid first binds to ATP, forming an aminoacyl-AMP intermediate.
This happens outside the editing site in the synthetase's main active site.
Transfer to tRNA:
The enzyme then attaches the amino acid to the tRNA, forming aminoacyl-tRNA.
This is where errors can happen, especially with similar-sized amino acids.
Editing step (Error Checking 🔍):
The enzyme then forces the newly attached amino acid into its editing pocket.
If it's incorrect (too small or slightly different in shape), it fits into the editing site, where it gets hydrolyzed and removed.
If it's correct, it won’t fit into the editing site, so it stays attached to tRNA and moves on to translation.
Multiple checkpoints: Most synthetases don’t rely on just one feature. They double-check several parts of the tRNA at once (anticodon + acceptor stem + other parts) to be extra sure. Think of it like verifying someone's face, fingerprint, and voice simultaneously.
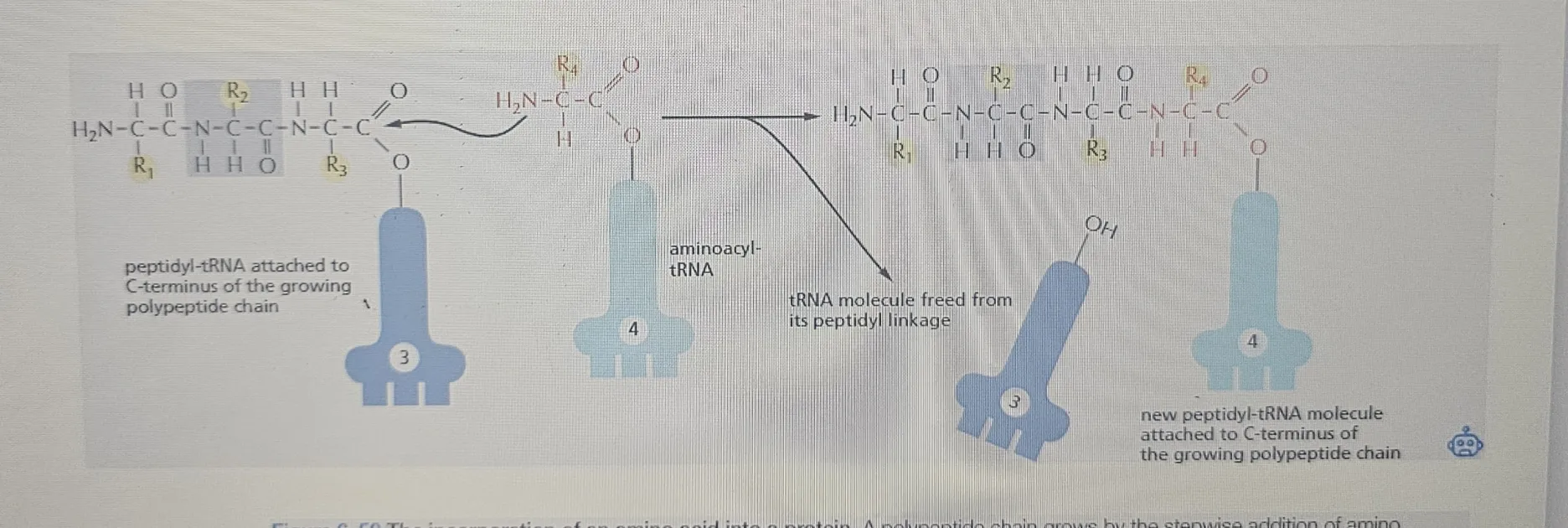
Amino Acids Are Added to the C-terminal End of a Growing Polypeptide Chain
After amino acids get matched to their correct tRNAs, it's go time for protein building:
The Main Reaction!
A protein is built by linking amino acids together with peptide bonds.
The carboxyl group (-COOH) of the growing chain meets the amino group (-NH₂) of the incoming amino acid, and then a peptide bond forms!
Which direction does the protein grow?
Always from the N-terminal (start) ➡ C-terminal (end).
Think of the N-terminal as the "front door" and the C-terminal as the "back door" of the protein house. 🏠
Where's the energy coming from?
The growing chain is always attached to a tRNA (called peptidyl-tRNA) through a high-energy covalent bond reaction. This bond provides the energy necessary for adding new amino acids during the elongation phase of translation.
Adding a new amino acid breaks that high-energy covalent linkage (which releases energy) but immediately forms a new identical high-energy bond on the next amino acid.
So the "activation energy" keeps passing down the line like a 🔥 hot potato!
Breaking the high-energy bond: When the amino acid is added to the tRNA, it involves breaking a high-energy covalent bond (between the amino acid and ATP). This release of energy is essential for the process because it provides the initial activation energy for aminoacylation (attaching the amino acid to the tRNA).
Absorbing the energy: The energy released from breaking that bond doesn't just disappear. Instead, it is immediately used to form a new high-energy bond between the amino acid and the tRNA. So, while the bond is broken, the energy is efficiently reused to continue the process.
Why is this clever?
Instead of each amino acid needing its energy to get added, the chain grows by "borrowing" the energy from the last linkage. This is called "head growth" 🧢:
The front (head) of the chain stays active.
Each new addition resets the high-energy situation, so the party keeps going
ATP hydrolysis = during tRNA charging (before the ribosome).
Peptide bond formation = powered by the high-energy bond from the charged tRNA, not fresh ATP.
Aminoacyl-tRNA Synthetases → Amino acid → Ribosome → Polypeptide
Step-by-step breakdown of where the energy comes from in protein synthesis:
1. ATP shows up EARLY — during tRNA charging
Before we even start making a protein, each amino acid has to get "loaded" onto its matching tRNA.This happens with the help of aminoacyl-tRNA synthetase, which uses ATP to make this happen.
Here’s what goes down:
ATP is used to attach the amino acid to its tRNA.
This creates a high-energy bond (called an ester bond) between the amino acid and the tRNA.
That bond stores the energy we’ll need later to link amino acids together.
1.5 So at this stage: Amino Acid + tRNA + ATP → Aminoacyl-tRNA (charged tRNA) + AMP + PPi
2. Ribosome
When the aminoacyl-tRNA arrives at the ribosome to join the growing protein chain. The amino acid still carries that high-energy bond from when ATP was spent during charging.
3. Peptide bond time!
At the ribosome, we link amino acids together to build the protein chain, and here's the genius part:
💥 The energy for the peptide bond (the bond that links amino acids) comes directly from breaking that high-energy bond between the amino acid and its tRNA.
👉 So, as the amino acid breaks free from its tRNA, that released energy is immediately used to form a new peptide bond between:
The carboxyl group (-COOH) of the growing chain
And the amino group (-NH₂) of the incoming amino acid.
4. Amino Acid
The chain is now attached to the next tRNA (the most recently arrived one).
The high-energy situation resets — because now the chain is linked to another tRNA through another high-energy bond.
This means the process keeps fueling itself without needing fresh ATP for every peptide bond.
Process happens simultaneously as the ribosome moves along the mRNA, facilitating the addition of amino acids to the growing polypeptide chain.
Aminoacyl-tRNA Synthetases are working constantly in the cytoplasm (or in the nucleus for some cases), attaching amino acids to tRNAs. As soon as an amino acid is attached to the tRNA, the tRNA is now "charged" and ready to bring its amino acid to the ribosome.
[ Amino Acid + tRNA ] --(ATP used)--> [ Charged Aminoacyl-tRNA (💥) ]
hydrolysis AMP 2PPI
Ribosomes are simultaneously working on the mRNA, reading the codons, and assembling the corresponding amino acids into a polypeptide chain. The ribosome needs the charged tRNA (from the synthetase) to continue building the protein.
At ribosome:
[ Growing Chain - tRNA ] + [ New Aminoacyl-tRNA (💥) ]
⬇
Peptide bond forms using 💥 from new tRNA
Chain grows → energy passes → repeat!
Ribosome Structure
Overview: So, proteins are assembled in the cytoplasm by ribosomes, but if they need to be processed (e.g., for secretion or membrane insertion), this happens in the rough ER. Translation uses the mRNA (which was made from DNA through transcription) to make proteins!
Transcription: DNA → mRNA (in the nucleus)
Transcription takes the DNA and creates a copy of it in the form of mRNA
Translation: mRNA → Protein (in the cytoplasm or rough ER)
Rough ER: Where some of the newly synthesized proteins are processed and modified for secretion or membrane insertion.
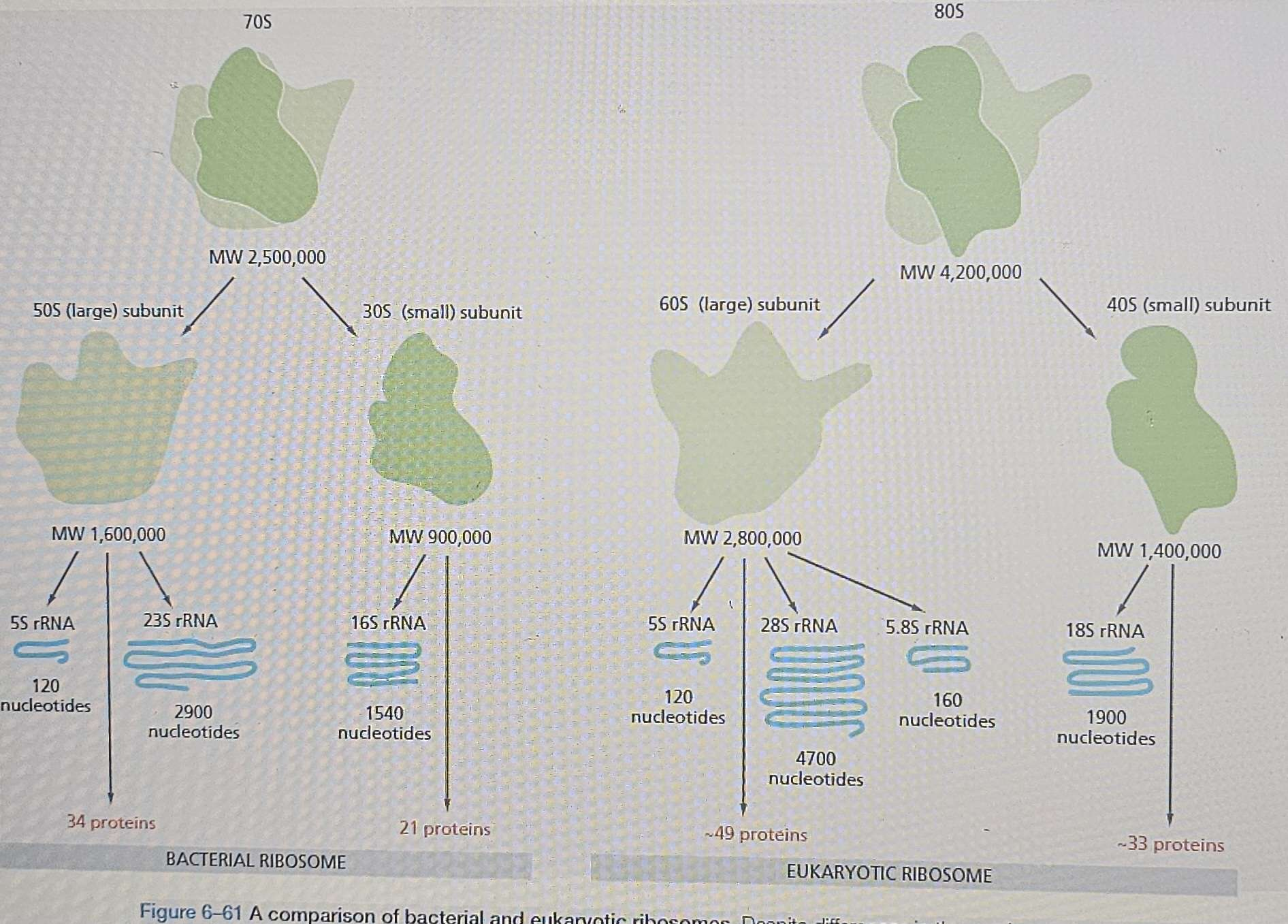
Composition: Large ribonucleoprotein complex ( form in the nucleolus), about 1 mistake every 10,000 amino acids, protein synthesis is performed in the ribosome consisting of two main subunits (small and large), several rRNA molecules, and over 50 proteins different proteins (the ribosomal proteins) and several RNA molecules, the ribosomal RNAs (rRNAs).
Assembly: Eukaryotic ribosomal subunits large and small are assembled in the nucleolus.
Quantity: Millions of ribosomes exist per eukaryotic cell.
Export:
Pre-rRNA Transcription (Inside the Nucleolus)
The main ribosomal RNA (rRNA) components are transcribed as pre-rRNA inside the nucleolus (a specialized region of the nucleus).
These pre-rRNAs then get processed into their mature forms (like 18S, 28S, and 5.8S rRNAs, depending on the subunit).
It starts as pre-rRNA (immature, unprocessed form).
It ends as rRNA (mature, functional form after processing).
Ribosomal Proteins (Synthesized in the Cytoplasm!)
Meanwhile, ribosomal proteins (which are needed for ribosome assembly) are made in the cytoplasm by existing ribosomes.
When the ribosomal subunits are in the cytoplasm, it means they have been exported out of the nucleus.
After they’re synthesized, they are imported back into the nucleus and enter the nucleolus. 🚀
Assembly of Ribosomal Subunits (Inside the Nucleus)
The rRNA and ribosomal proteins assemble into two separate ribosomal subunits (small and large) inside the nucleolus.
Export to the Cytoplasm
Once assembled, the two ribosomal subunits are individually exported from the nucleus to the cytoplasm. 🏃♂💨
They stay separate until it’s time to start translating mRNA.
Final Assembly & Protein Synthesis (In the Cytoplasm!)
When protein synthesis begins, the small subunit binds to mRNA first.
Then, the large subunit joins to form a fully functional ribosome.
The ribosome can now synthesize proteins by linking amino acids together. 🔗💥
Ribosome Organization:
Small subunit binds to mRNA first:
The small ribosomal subunit is the first to interact with the mRNA strand. It reads the mRNA and starts the process of translation(tRNA) binding. Acts as the framework on which the tRNAs are accurately matched to the codons of the mRNA
The small subunit is involved in reading the mRNA and bringing in the tRNAs.
Large subunit binds to the small subunit:
Once the small subunit is attached to the mRNA, the large ribosomal subunit then binds to the small subunit to form a functional ribosome. This is when the ribosome is fully assembled and ready to start protein synthesis. The large subunit catalyzes the formation of the peptide bonds that link the amino acids together into a polypeptide chain.
The polypeptide chain is the protein being synthesized.
The large subunit is involved in making the peptide bonds between the amino acids in the growing polypeptide chain.
Summary:
In the nucleus, pre-rRNA is transcribed from DNA and processed into mature rRNA. At the same time, ribosomal proteins are synthesized in the cytoplasm and exported to the nucleus, where they enter the nucleolus. Inside the nucleolus, rRNA and ribosomal proteins are assembled separately into the small and large ribosomal subunits.
Once assembled, these ribosomal subunits are exported individually from the nucleus to the cytoplasm. In the cytoplasm, protein synthesis begins when the small subunit binds to mRNA. The large subunit then joins the small subunit, forming a functional ribosome. The ribosome can now synthesize proteins by linking amino acids together based on the mRNA code.

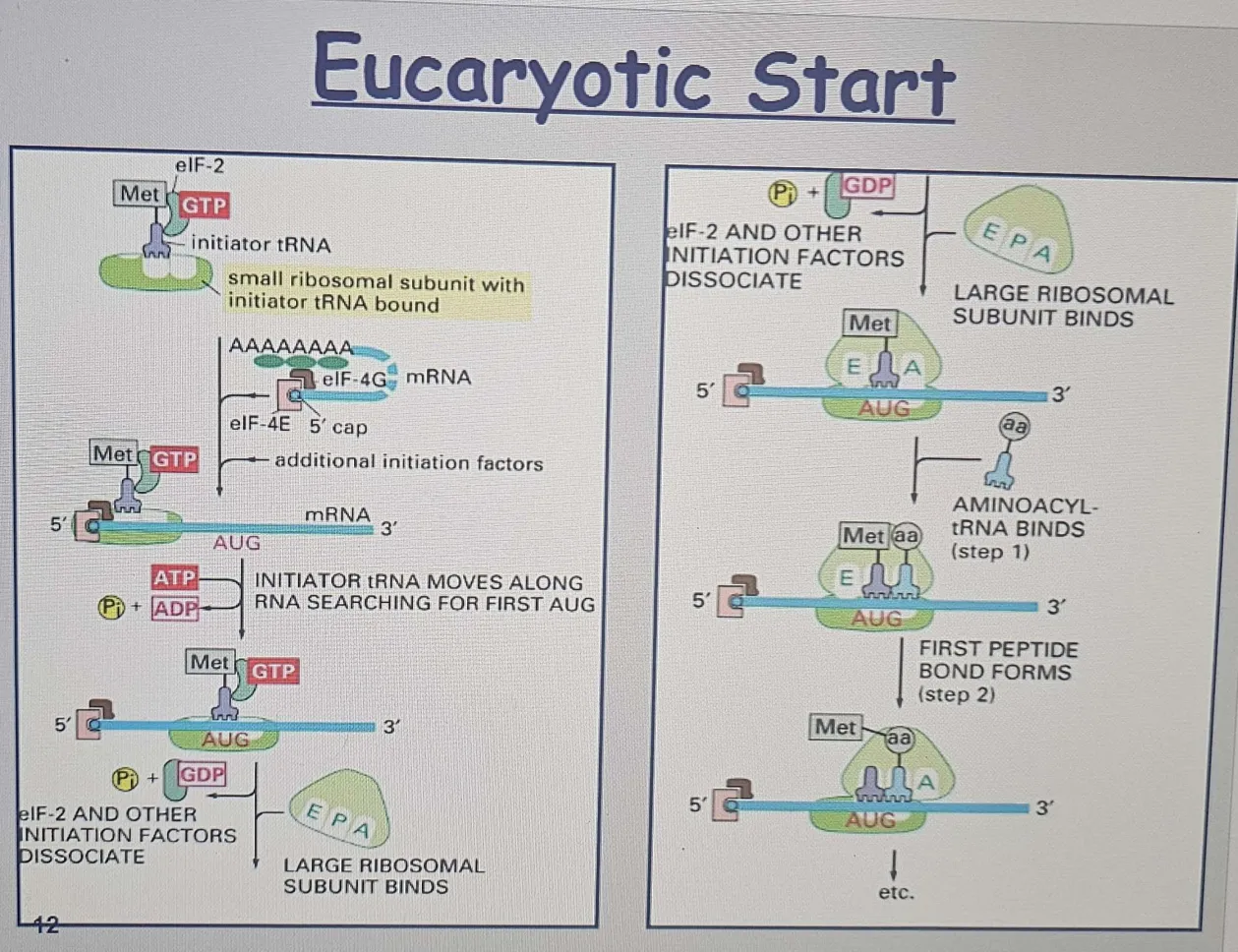
Eukaryotic Initiation
This phase happens before elongation and sets up the ribosome on the mRNA so translation can begin.
Importance of Initiation:
The site where protein synthesis begins is crucial as it determines the reading frame of the mRNA.
Error at this stage leads to the misreading of the entire sequence, producing a nonfunctional protein.
A. Initiator tRNA Loading
1. Initiator tRNA: The first tRNA (which carries methionine in eukaryotes) or formylmethionine (in bacteria) it binds to the small ribosomal subunit (SRS).
This tRNA binds directly to the small ribosomal subunit (40S) even before the full ribosome is assembled!
It goes straight to the P site (unlike most tRNAs that enter at the A site).
Of all the aminoacyl-tRNAs in the cell, only the methionine-charged initiator tRNA is capable of tightly binding the small ribosome subunit without the complete ribosome being present, and unlike other tRNAs it binds directly to the A site
This tRNA’s job? Find and recognize the start codon (AUG) on the mRNA.
B. Eukaryotic Initiation Factors (eIFs)
2. eIFs are helper proteins that bind to the small ribosomal subunit (SRS).
These initiation factors help the ribosome recognize the 5' end of the mRNA (the cap structure) and make the mRNA ready for translation.
It binds that have previously bound two initiation factors, eIF4E and eIF4G
C. Scanning for the Start Codon
3. ATP-powered helicase activity (eIFs) helps in moving along the mRNA.
This scanning process requires ATP hydrolysis (breaking down ATP for energy).
4. The small ribosomal subunit (SRS), along with the initiator tRNA and eIFs, starts scanning the mRNA from the 5' cap towards the first AUG codon (start codon).The ribosome scans until it finds the first suitable AUG to begin translation.
D. Large Subunit Joins
5. After the first AUG codon is found, eIFs dissociate, the large ribosomal subunit (60S) joins the small subunit (40S), forming the complete 80S ribosome.
Leaky Scanning: Not all AUGs are read; some may be ignored for later ones, Why? The surrounding sequence wasn’t ideal, so the ribosome decided to keep looking, allowing the synthesis of proteins with similar functions but different localization.
Scans for AUG: If the first AUG does not have the correct surrounding sequence, the ribosome may skip to the next AUG.
This produces multiple proteins from one mRNA, differing in their N-terminus.
Common in genes encoding proteins with or without a signal sequence so that the protein is directed to two different compartments in the cell.
6. The initiator tRNA stays in the P site, and the A site is vacant. Now, the ribosome is fully assembled, and the translation process can begin.
Bacterial Translation Initiation
A. Lack of 5' Cap and Consensus Sequences
No 5' Cap: Unlike eukaryotic mRNA, bacterial mRNA lacks a 5' cap structure.
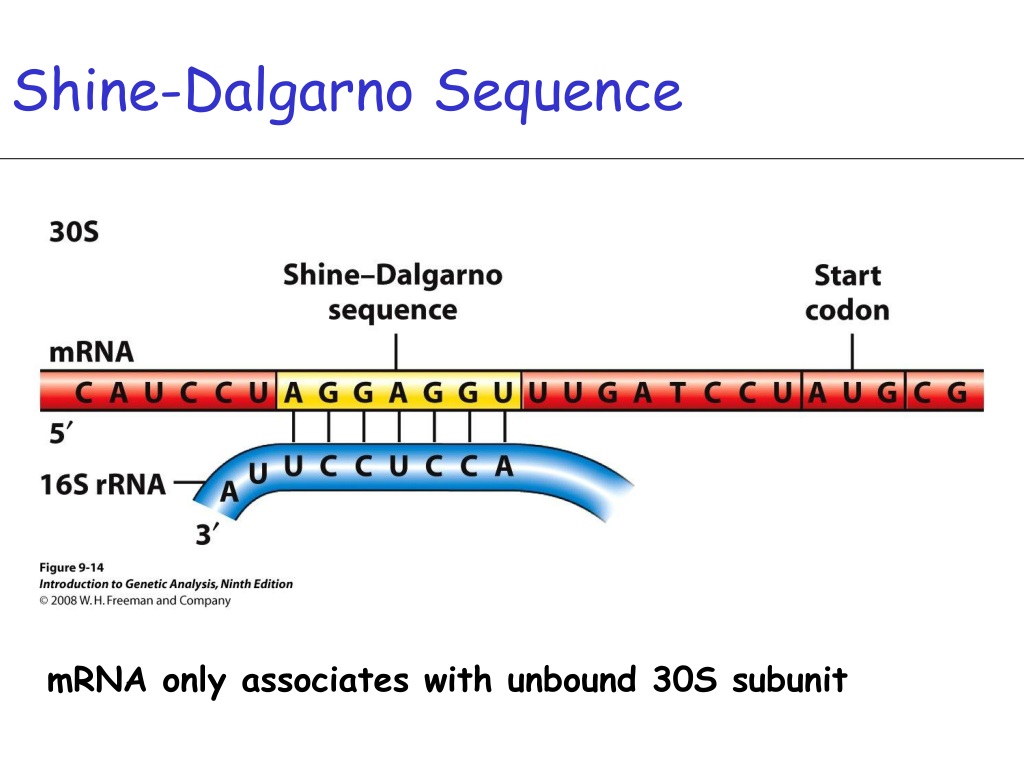
Shine-Dalgarno Sequence: Instead of the 5' cap, bacterial mRNA has a Shine-Dalgarno sequence, a consensus sequence found just upstream of the AUG codon. This sequence helps the small ribosomal subunit (30S) bind to the mRNA and start scanning for the AUG codon.
Not randomly in the middle of the entire mRNA.
Location: About 5-9 nucleotides upstream of the start codon (AUG).
It’s located just before the AUG start codon in the 5' untranslated region (5' UTR).
B. Polycistronic mRNA
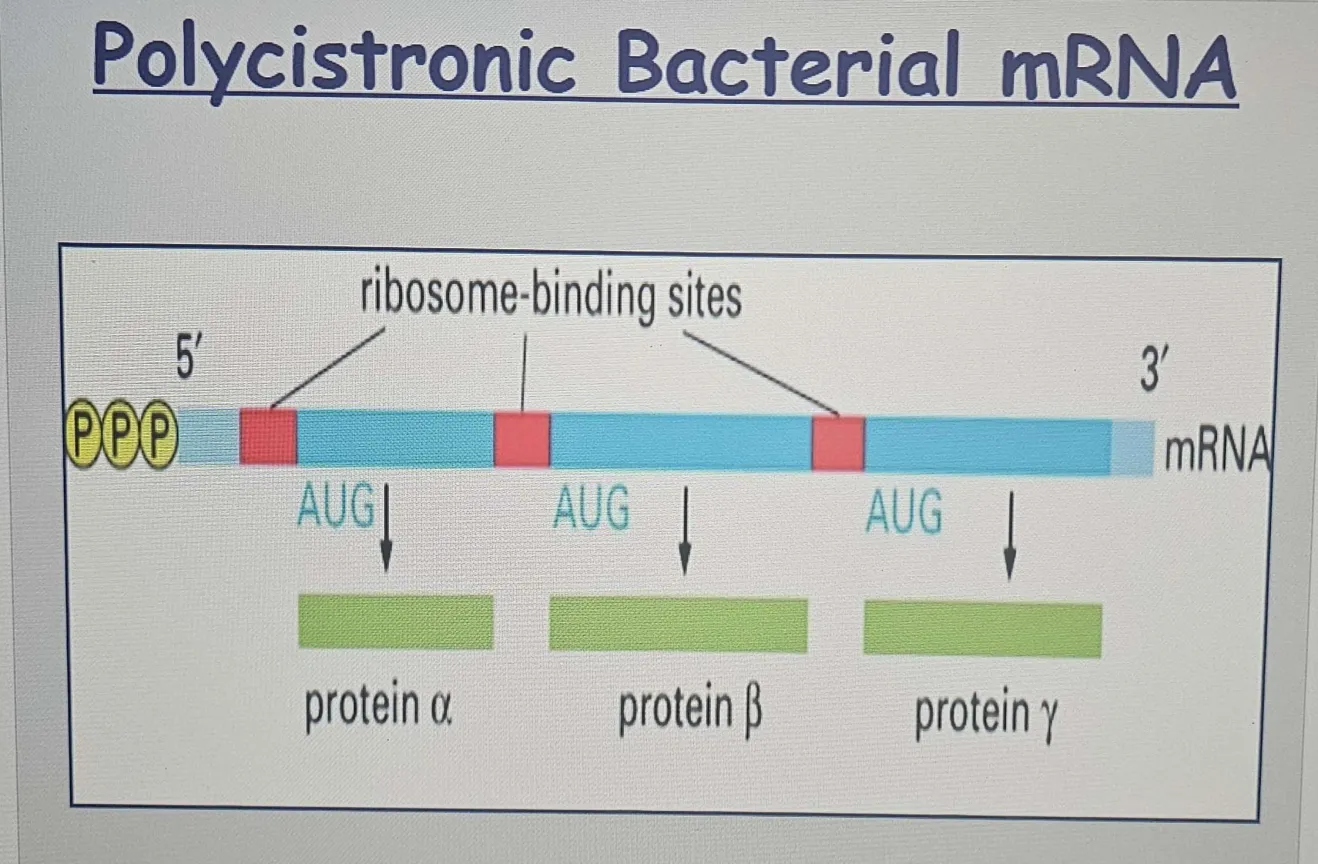
Polycistronic mRNA: Bacterial mRNA is often polycistronic, meaning it can contain multiple coding regions (or cistrons) that each encode for a different protein. This allows bacteria to efficiently synthesize multiple proteins from a single mRNA molecule.
The presence of multiple Shine-Dalgarno sequences allows for separate translation initiation events at different AUG codons, enabling the synthesis of several proteins from the same mRNA.Since each Shine-Dalgarno (SD) sequence is located upstream of an AUG start codon, there can be multiple independent starting points for translation on the same mRNA.
C. Ribosome Binding and Scanning
The small ribosomal subunit (30S) binds to the Shine-Dalgarno sequence and scans for the first AUG codon to start translation, similar to the scanning process in eukaryotes but without the 5' cap recognition.
D. Large Subunit Joins
After the AUG codon is found, the large ribosomal subunit (50S) binds to the small subunit (30S), forming the 70S ribosome, and translation begins.
Key Differences:
5' Cap: Only eukaryotic mRNA has a 5' cap.
Shine-Dalgarno vs. 5' Cap: Bacteria use the Shine-Dalgarno sequence to help ribosomes bind, while eukaryotes use the 5' cap.
Polycistronic vs. Monocistronic: Bacteria can produce multiple proteins from one mRNA (polycistronic), while eukaryotes produce a single protein per mRNA (monocistronic).
Initiation Mechanism: The way ribosomes initiate translation (based on the Shine-Dalgarno sequence in bacteria vs. 5' cap scanning in eukaryotes) differs.
Ribosome Activity During Protein Synthesis (Elongation Phase)
Once initiation is complete, the ribosome is ready for elongation, which is the actual synthesis of the protein. Here's what happens during elongation:
🔵 mRNA and Ribosome Interaction
The small ribosomal subunit binds to the mRNA, and the ribosome reads the mRNA codons.
🟢Elongation: tRNA Binding
Ribosome contains four binding sites for RNA molecules: one is for the
mRNA and three (called the A site, the P site, and the E site) are for tRNAs
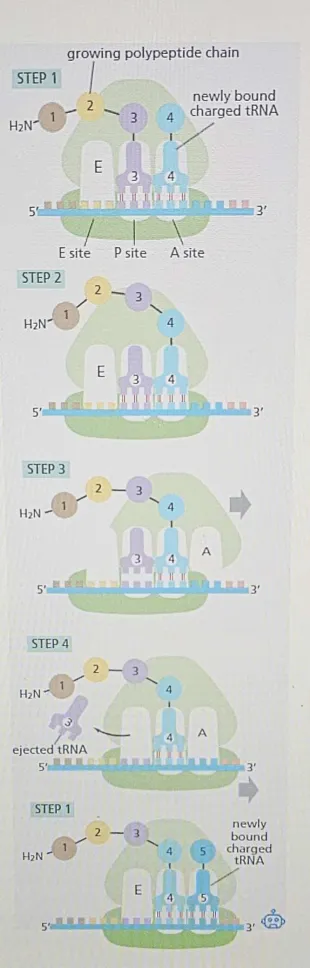
tRNA Binding (Step 1):
A new tRNA carrying the next amino acid enters the A site of the ribosome.
The anticodon of the tRNA pairs with the complementary codon of the mRNA that's positioned in the A site.
The A and P sites are close enough together for their two tRNA molecules to be forced to form base pairs with adjacent codons on the mRNA molecule. This feature of the ribosome maintains the correct reading frame on the mRNA.
The ribosome uses the wobble hypothesis to allow some flexibility in base pairing at the 3rd position of the codon-anticodon interaction. This flexibility enables the ribosome to accommodate variations in tRNA molecules, which can enhance the efficiency of protein synthesis.
A site: Accepts new tRNA with the correct amino acid. Incoming aminoacyl-tRNA enters the A-site; the peptide chain elongates. (aa)
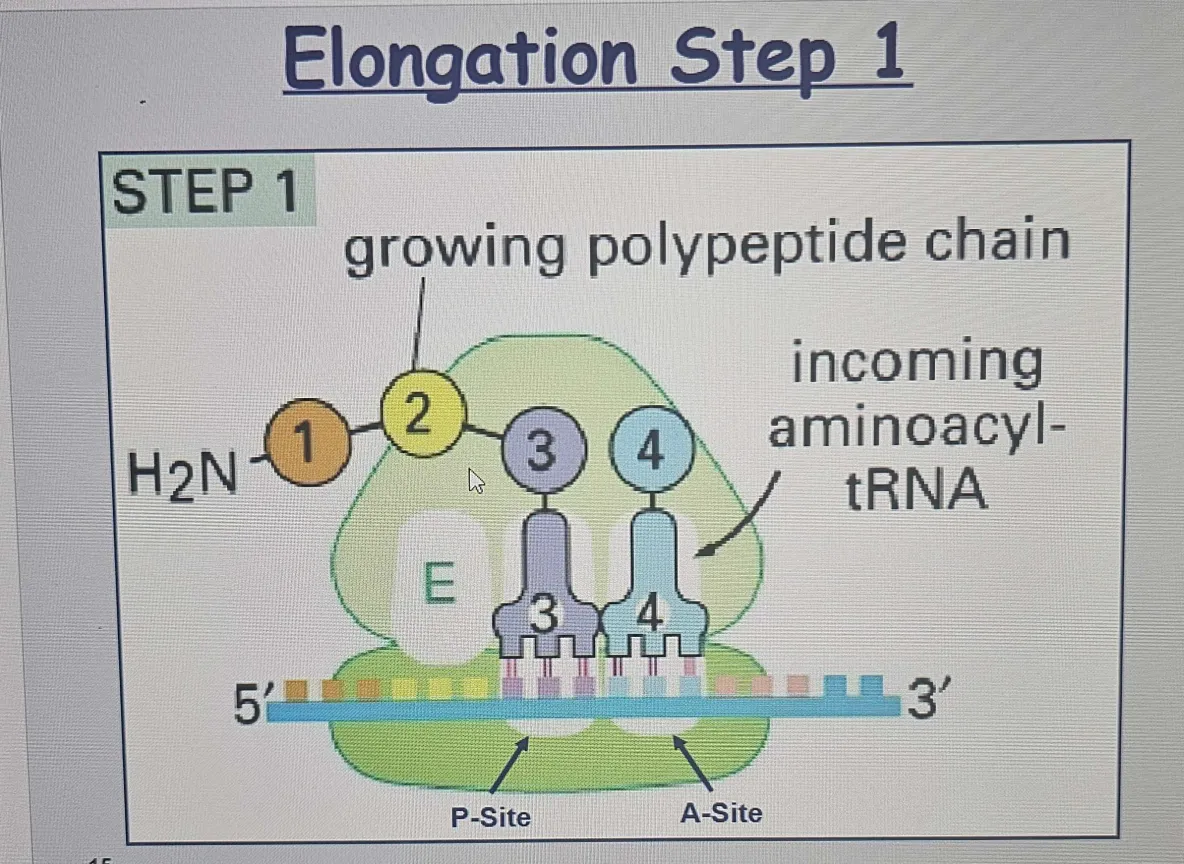
🟡 Peptide Bond Formation (Step 2):
P site: Holds the tRNA with the growing polypeptide chain.
Bond energy forms a new bond between amino acids; large subunit shifts.
Once the tRNA enters the A site, The peptidyl transferase activity of the ribosome catalyzes the formation of a peptide bond between the carboxyl group of the amino acid on the P-site tRNA and the amino group of the new amino acid on the A-site tRNA.
This is not about holding things in place by the carboxyl group—it’s about forming a new peptide bond between the two amino acids. The carboxyl group of the growing chain (on the P-site tRNA) reacts with the amino group of the incoming amino acid (on the A-site tRNA), creating the bond that holds the new amino acid to the growing chain.
The amino acid in the P site is transferred to the tRNA in the A site,it reacts with the amino group of the new amino acid from the A site, and the result is the formation of the peptide bond that links the two amino acids together.
When the carboxyl group from the polypeptide chain (in the P site) reacts with the amino group of the incoming amino acid (in the A site), the carboxyl group is indeed "broken" or, more specifically, it loses the hydroxyl group (OH) as part of the dehydration reaction.
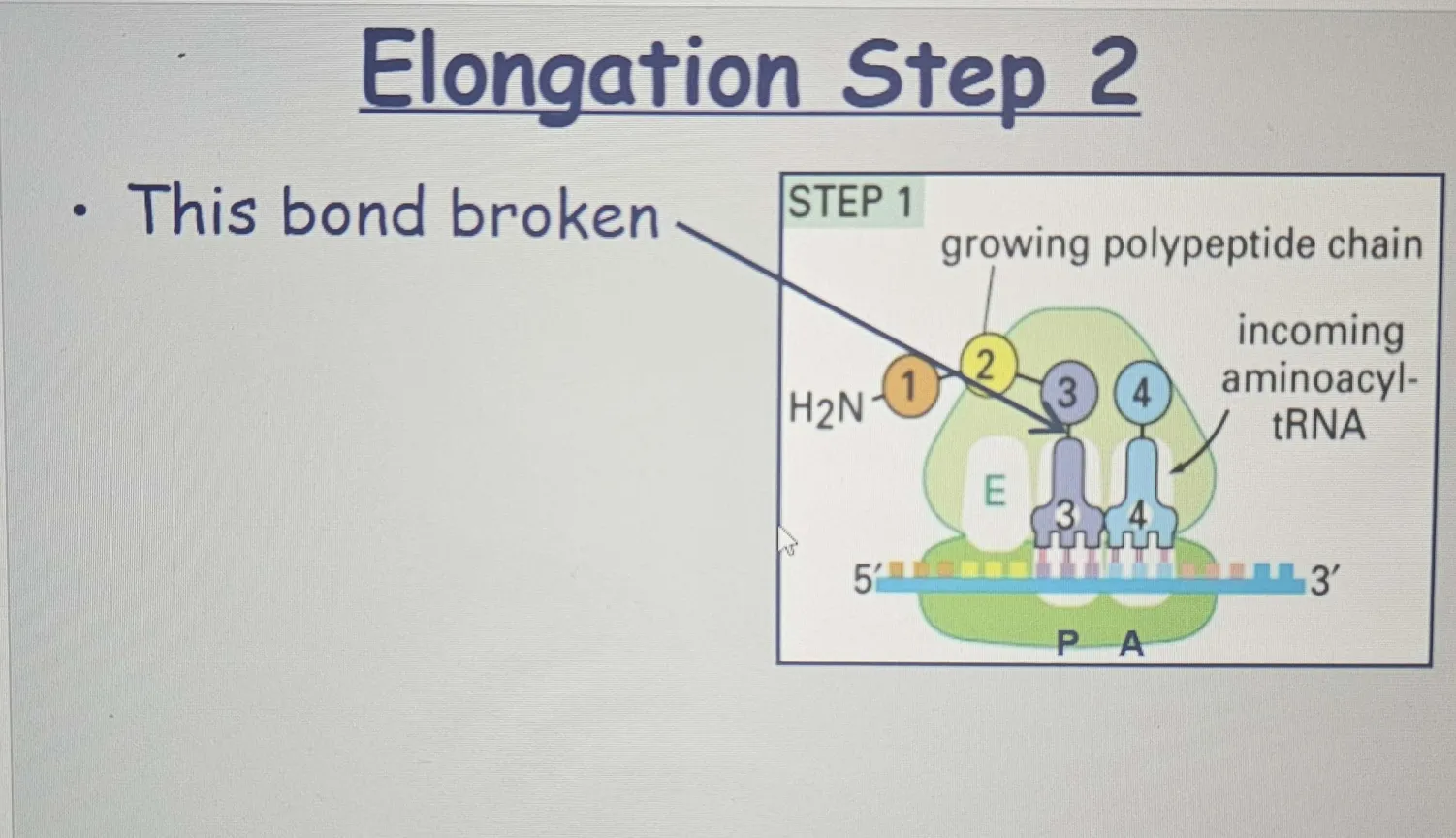

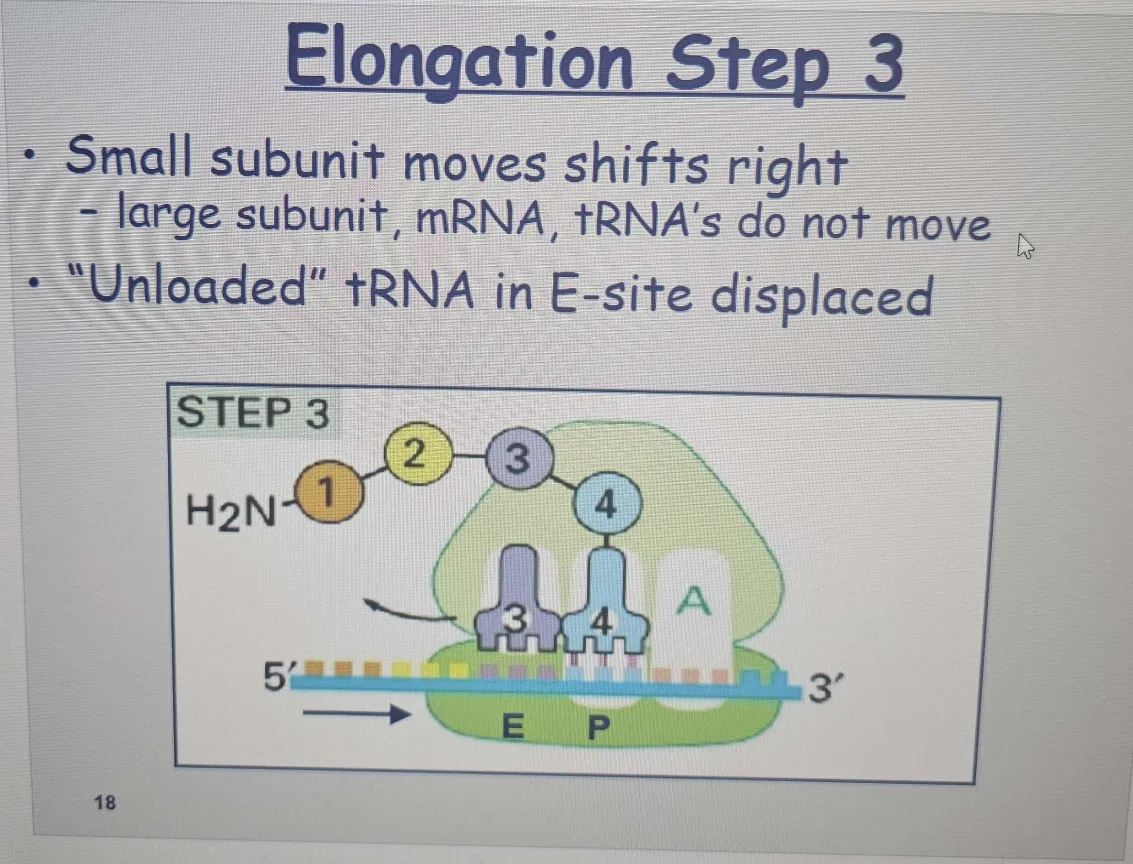
🟠 Large Subunit Translocation (Step 3):
After the peptide bond is formed, the large ribosomal subunit moves along the mRNA by three nucleotides
This movement shifts the tRNA in the A site to the P site, and the tRNA in the P site moves to the E site before exiting.
The tRNA brings in the appropriate amino acid based on the mRNA codon.
E site: After the peptide bond is formed, the tRNA that was carrying the growing polypeptide chain moves from the P site to the E site (exit site) on the ribosome.This happens because the polypeptide is now attached to the tRNA in the A site, and the growing chain is ready to continue being synthesized.
🟠 Small Subunit Translocation (Step 4):
Following the large subunit translocation, another series of conformational changes moves the small subunit and its bound mRNA exactly three nucleotides, ejecting the spent tRNA from the E site and resetting the ribosome so it is ready to receive the next aminoacyl-tRNA.
This keeps the ribosome in sync, ensuring the correct positioning of the A site, P site, and E site.
After this movement, the A site is empty and ready to accept a new tRNA, and the cycle can begin again with the next amino acid.
The Blob
Small Subunit (on the bottom):
Function: It binds to the mRNA and decodes the genetic information. It helps ensure that the right tRNA is matched with the correct codon on the mRNA.
Position: The small subunit is what initially interacts with the mRNA and helps align the mRNA with the tRNA.
Large Subunit (on the top):
Function: It contains the peptidyl transferase activity, which catalyzes the formation of peptide bonds between the amino acids brought by the tRNAs. This is where the polypeptide chain is actually formed.
Position: The large subunit sits on top and facilitates the process of protein synthesis by joining the amino acids together.
🟣Repeat
Step 1 is then repeated with a new incoming aminoacyl-tRNA, and so on.
The process repeats, with new tRNAs entering the A site, adding their amino acids, and forming the polypeptide chain as the chain grows from its amino to its carboxyl end.
This reaction frees the carboxyl end of the growing polypeptide chain from its attachment to a tRNA molecule, and since only this attachment normally holds the growing polypeptide to the ribosome, the completed protein chain is immediately released into the cytoplasm. The ribosome releases the finished protein and dissociates into its small and large subunits to start the process over for the next mRNA. The ribosome then releases its bound mRNA molecule and separates into the large and small
subunits. These subunits can then assemble on this or another mRNA molecule to begin a new round of protein synthesis.
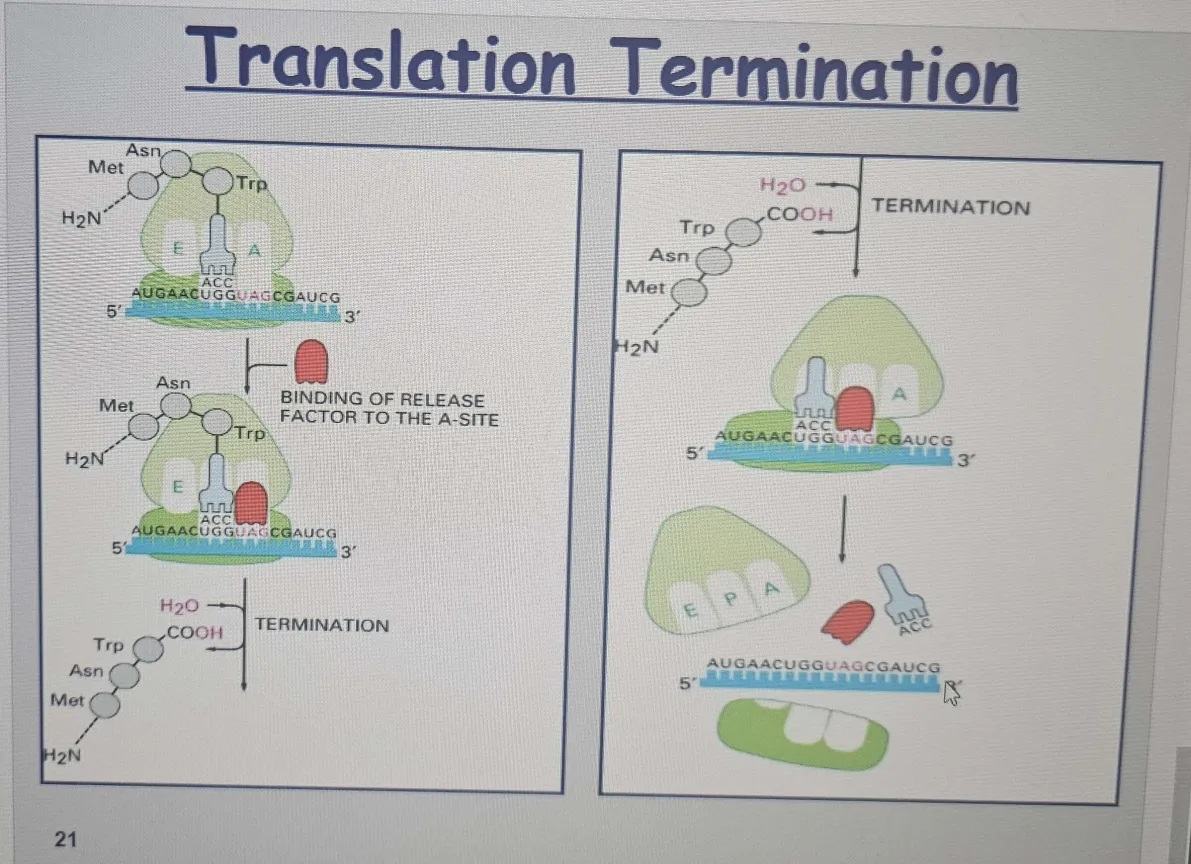
🔴 Stop Codon Encounter and Termination
When the ribosome reaches a stop codon (UAA, UAG, or UGA), it signals the end of translation.
Release Factor (RF): Mimics tRNA and adds a water molecule instead of an amino acid, releasing polypeptide.
No tRNA in the A site: The A site is empty when a stop codon is reached, so no tRNA enters to add another amino acid.
Release factor: A release factor (a protein, not a tRNA) binds to the A site instead of a tRNA. This release factor causes the ribosome to catalyze a reaction that adds water (H₂O) to the end of the polypeptide chain instead of an amino acid.
Breaking the bond: This water molecule breaks the bond between the carboxyl group of the polypeptide and the tRNA in the P site, which releases the completed protein from the ribosome
Role of Elongation Factors
Involvement of GTP; incorrect pairing results in preferential dissociation of mismatched tRNAs.
1. Role of Elongation Factors (EFs)
Two elongation factors assist translation by entering and leaving the ribosome during each cycle:
Bacteria: EF-Tu and EF-G
Eukaryotes: EF1 and EF2
These factors hydrolyze GTP to GDP, undergoing conformational changes that drive translation forward.
Without these factors, protein synthesis is slow, inefficient, and inaccurate.
GTP hydrolysis ensures translation occurs in the "forward" direction, increasing efficiency.
2. EF-Tu and Translation Accuracy (codons)
EF-Tu binds GTP and aminoacyl-tRNAs, helping position the tRNA in the A site.
The 16S rRNA (small ribosomal subunit) checks the codon-anticodon match:
A correct match ( Increases accuracy) triggers a conformational change in the ribosome → GTP hydrolysis by EF-Tu → tRNA is released for use.
An incorrect match fails to trigger this change → the incorrect tRNA falls off before being used.
This means that while a correct codon-anticodon match forms a stronger bond than an incorrect one due to favorable free-energy changes, the difference in binding strength alone is not enough to guarantee high accuracy in translation.
In other words, an incorrect tRNA could still bind (just not as tightly), which means the ribosome needs additional mechanisms (like elongation factor EF-Tu and ribosomal proofreading) to ensure that only the correct tRNA stays and is used in protein synthesis.
3. Two Proofreading Steps for Accuracy
1. EF-Tu Check: Ensures only correctly bound tRNAs undergo GTP hydrolysis and are released for translation.
Protein folding: First, the 16s rRNA in the small subunit of
the ribosome assesses the “correctness” of the codon-anticodon match by folding around it and probing its molecular details
Correct match is found, the rRNA closes tightly around the codon-anticodon pair, causing a conformational change in the ribosome that triggers GTP hydrolysis by EF-Tu. Only when GTP is hydrolyzed does EF-Tu release its grip on the aminoacyl-tRNA and allow it to be used in protein synthesis.
The anticodon on the tRNA correctly pairs with the codon on the mRNA in the A site of the ribosome.
This correct match causes a conformational change in the ribosome.
The ribosome triggers EF-Tu to hydrolyze GTP to GDP.
When GTP is hydrolyzed to GDP, EF-Tu releases the aminoacyl-tRNA (tRNA with its amino acid) and allows the tRNA to be used in the peptidyl transferase reaction (forming the peptide bond).
Incorrect codon–anticodon matches do not readily trigger this conformational change, and these errant tRNAs mostly fall off the ribosome before they can be used in protein synthesis.
The anticodon on the tRNA does not correctly pair with the codon on the mRNA.
Since the match is incorrect, the ribosome does not undergo the conformational change.
EF-Tu does not hydrolyze GTP because the tRNA is not properly matched. So, GTP remains bound to EF-Tu, and EF-Tu does not release the tRNA.
The incorrect tRNA is rejected and leaves the ribosome without participating in the peptide bond formation.
If the match is correct, GTP is hydrolyzed, and the tRNA is allowed to participate in protein synthesis.
If the match is incorrect, GTP is NOT hydrolyzed, and the tRNA is ejected from the ribosome without contributing to protein synthesis.
2. Ribosome Delay Check:
Delay: There is a short time delay as the amino acid
carried by the tRNA moves into position on the ribosome.
This time delay is shorter for correct than incorrect codon-anticodon pairs. Thus, the correct codon-anticodon pairs move efficiently into position.
Incorrect pairs experience a delay and dissociate faster because their interaction/bond with the codon is weaker, preventing incorrect incorporation.Even some correctly matched tRNAs may fall off, but most incorrect ones are removed before they are used in translation.
This double-check mechanism ensures 99.99% accuracy in translation.
4. Final Error Detection Mechanism
If an incorrect amino acid still gets incorporated, the incorrect codon‒
anticodon interaction in the P site of the ribosome (which would occur after the misincorporation) causes an increased rate of future misreading in the A site.
The incorrect codon-anticodon interaction in the P site disrupts the ribosome’s ability to accurately read the next codon in the A site.This leads to more mistakes, making it likely that more incorrect amino acids will be added.
This leads to premature translation termination by release factors, preventing the creation of faulty proteins.
If too many errors accumulate, the ribosome stops translation early by triggering release factors. Normally, these release factors act when the translation of a protein is complete; here, they act early.
The misfolded protein is degraded, stopping further waste of resources.
which terminates protein synthesis and marks the faulty protein for degradation.

Overcoming Limitations of Complementary Base-Pairing
Complementary Base-Pairing Accuracy
DNA replication, repair, transcription, and translation all rely on base-pairing rules (G≡C, A=T/U).
However, base-pairing alone only provides a 10- to 100-fold difference in affinity between correct and incorrect matches. This alone cannot explain the extremely high accuracy in DNA replication, transcription, and translation.To achieve much greater accuracy, cells use additional mechanisms beyond simple base-pairing. In the case of the ribosome, two key principles help:
Induced Fit (First Accuracy Check) # 3:1
The ribosome folds around the codon-anticodon pair.
This folding only completes if the match is correct, allowing translation to proceed.
If the match is incorrect, the folding fails, and the tRNA will likely dissociate.
A similar mechanism is seen in RNA polymerase during transcription.
Kinetic Proofreading (Second Accuracy Check) # 3:2
After codon-anticodon pairing, GTP is hydrolyzed, making the step irreversible.
A time delay follows before the aminoacyl-tRNA is used in peptide bond formation.
Incorrect matches are more likely to dissociate because:
Weaker interaction with the codon, i.e wrong tRNA with the codon is weaker
Longer delay for incorrect matches, increasing the chance they fall off.
This extra proofreading step ensures greater accuracy than base-pairing alone.
Kinetic proofreading consumes ATP/GTP, making the translation more energy-intensive, but it ensures 99.99% accuracy in protein synthesis.
Analogy:
A worker (enzyme) grabs a piece (molecule) and checks if it fits.
If it fits well, the worker secures it quickly and moves on. ✅
If it doesn’t fit quite right, the worker holds it for a bit longer, double-checks, and most likely throws it out before moving on.
This extra "checking time" makes sure only the right pieces stay, preventing errors from slipping through!
In the analogy, you're using the worker’s pause or hesitation, which allows them to decide based on closer inspection. If something doesn't fit, they hesitate, double-check, and discard it. In this sense, waiting allows the worker to "release" or discard the item after carefully considering whether it’s correct or not. So, “waiting” here is about giving that extra moment to ensure the final decision—whether to keep or discard—has the right judgment. It’s not really about the time itself causing the release, but rather that the pause allows for a better decision (in the same way, kinetic proofreading pauses for error-checking before continuing).
Energetic Cost of Translation
Accuracy vs. Speed: translation requires a compromise between accuracy and speed. To maintain accuracy (one mistake per 10,000 amino acids), there need to be time delays during translation, which slow down the process. In bacteria, the translation rate is typically around 20 amino acids per second, but this speed is compromised by accuracy.
Ribosomes operate with remarkable efficiency: in one second, a eukaryotic ribosome adds 2 amino acids to a polypeptide chain; the ribosomes of bacterial cells operate even faster, at a rate of about 20 amino acids per second
Mutants and Trade-off: Mutant bacteria with a more accurate ribosome (due to longer delays between amino acid additions) are much slower in protein synthesis. These mutants have higher translation accuracy, but their slow rate means they can barely survive.
Energy Cost of Accuracy: Ensuring accurate protein synthesis consumes much free energy. Accuracy > OVER speed
At least high energy bonds (ATP or GTP) per peptide bond
highest per bond cost of any synthesis reaction
For every new peptide bond formed, four high-energy phosphate bonds are used:
Two are used for charging the tRNA with an amino acid.
Two more drive the cycle of elongation during protein synthesis.
Additionally, energy is required to remove incorrect amino acids and reject incorrect tRNAs through proofreading mechanisms.
Proofreading and Energy Consumption:
Proofreading (ensuring the correct amino acid is added) consumes even more energy than initially expected.
The proofreading mechanisms need to remove some correct interactions as well to maintain the high accuracy, which increases the energy cost.
Polyribosomes (Polysomes)
Function: Multiple ribosomes translate the same mRNA concurrently, enhancing protein synthesis rates.
Polyribosomes (Polysomes) and Protein Synthesis
Polyribosomes (Polysomes):
Multiple ribosomes can translate the same mRNA simultaneously, forming large cytoplasmic assemblies.
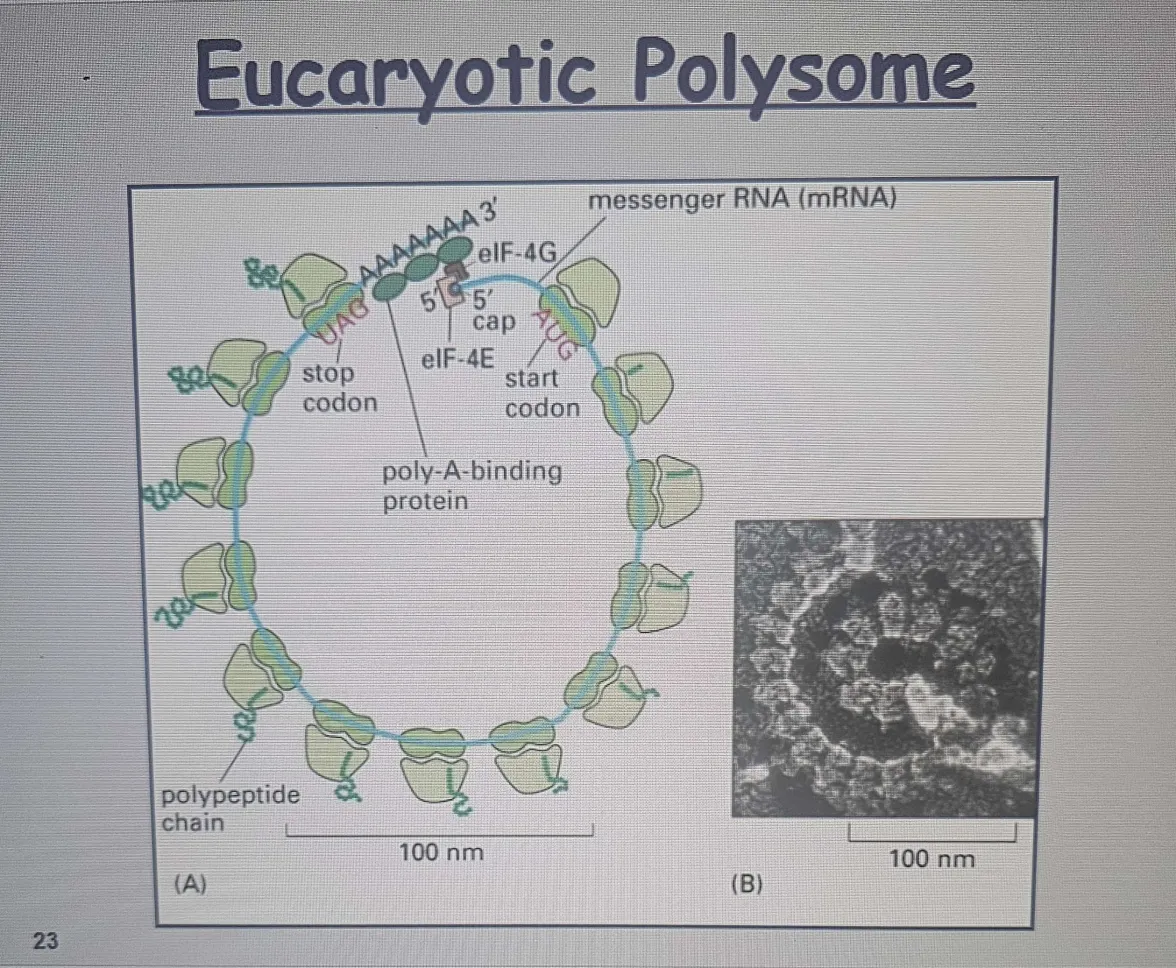
Spacing: Ribosomes are typically 80 nucleotides apart on a single mRNA.
Multiple initiations occur on each mRNA, allowing efficient protein synthesis.
Importance of Polysomes:
Multiple ribosomes on a single mRNA allow the cell to produce more protein molecules in a given time than if only one ribosome translated the mRNA.
Bacterial Translation:
Bacteria don't require mRNA processing and are accessible to ribosomes while it is being made. Ribosomes attach to the free end of bacterial mRNA and start translating while the RNA is still being transcribed, even before the transcription of that RNA is complete, following closely behind the RNA polymerase as it moves along DNA.
This enables them to rapidly respond to environmental changes and efficiently synthesize proteins as needed.
In bacteria, since there’s no nucleus, ribosomes can start translation while transcription is still happening.
Eukaryotic Translation:
In eukaryotes, as we have seen, the 5ʹ and 3ʹ ends of the mRNA interact therefore, as soon as a ribosome dissociates, its two subunits are in an optimal position to reinitiate translation on the same mRNA molecule.
In eukaryotes, mRNA must be fully transcribed, processed (capping, splicing, polyadenylation), and form in the cytoplasm; exported from the nucleus before ribosomes can bind and start translation.
In bacteria, since there’s no nucleus, ribosomes can start translation while transcription is still happening.
Process Differences: Bacterial polysomes can begin synthesis before transcription completion, a capability not found in eukaryotes.
Exceptions to the Genetic Code:
Most organisms follow the standard genetic code, but some exceptions exist.
Candida albicans translates CUG as serine, whereas most organisms translate it as leucine.
Mitochondria often have different codon assignments:
In mammalian mitochondria, AUA is translated as methionine, but in the cytosol, it codes for isoleucine.
Translation Recoding:
Some cells can modify codon interpretation based on additional mRNA sequence information.
This process allows the incorporation of non-standard amino acids into proteins.
Selenocysteine (21st Amino Acid):
Found in bacteria, archaea, and eukaryotes.
Contains selenium instead of sulfur (compared to cysteine).
Synthesized from serine attached to a special tRNA.
Inserted into proteins at UGA codons, which usually signal a stop.
Requires a specific mRNA sequence near the UGA codon to trigger this recoding event.
Eukaryotic polysome: A structure consisting of multiple ribosomes attached to a single mRNA, allowing for simultaneous translation of a protein in eukaryotic cell
Molecular Chaperones and Protein Folding
Protein folding
Folding Process: Occurs during synthesis; secondary and tertiary structures develop concurrently.
The Role of Molecular Chaperones
Most proteins do not fold correctly on their own during synthesis
Molecular chaperones assist in proper folding and prevent incorrect folding pathways.
Speed up/facilitate tertiary folding and repair minor misfolds.
Why are chaperones necessary?
Many folding pathways exist, but not all lead to the correct, stable structure.
Incorrect folding can cause proteins to become "kinetically trapped" in unstable structures.
Some misfolded proteins aggregate irreversibly, forming dead end of nonfunctional and toxic clumps (potentially dangerous) structures.
How Chaperones Recognize Misfolded Proteins
Misfolded proteins expose hydrophobic regions that should be buried inside the protein’s core.
These exposed hydrophobic surfaces tend to stick together →, forming dangerous aggregates.
Chaperones recognize these hydrophobic patches and bind to them using their own surfaces.
By doing this, chaperones prevent aggregation and help proteins refold correctly.
The Danger of Protein Aggregation
Protein aggregation can be toxic to cells, leading to diseases.
Many inherited diseases (like Alzheimer’s, Parkinson’s, and Huntington’s) involve protein misfolding and aggregation.
Chaperones help reduce disease risk by ensuring proper protein folding.
How Chaperones Work
Many molecular chaperones are called heat-shock proteins (designated, hsp), because they are synthesized dramatically after a brief exposure of cells to an elevated temperature.
This reflects the operation of a feedback system that responds to an increase in misfolded proteins (such as those produced by elevated temperatures) by boosting the synthesis of the chaperones that help these proteins
refold.
Chaperones bind to misfolded proteins and give them another chance to fold correctly.
They may use ATP-driven cycles to repeatedly bind and release proteins until proper folding.
There are several types of chaperones, each with specific mechanisms to aid protein folding.
In sum
Heat causes misfolding, and that same heat triggers a feedback response that increases chaperone (HSP) production to stop proteins from clumping.
They don’t generate heat but are produced in response to heat-induced stress.
Their job is to prevent misfolded proteins from clumping and help them refold.
Once they stabilize the proteins, the stress response decreases, reducing the need for more chaperones.
Heat (or other stress) = Fire 🔥
Misfolded proteins = Burning debris 🏚
Chaperones = Firefighters 🚒
They rush in when misfolded proteins appear.
They prevent the spread (stop aggregation).
They repair what they can (help refold proteins).
If a protein is too damaged, they remove the wreckage (send it for degradation).
Types of Molecular Chaperones
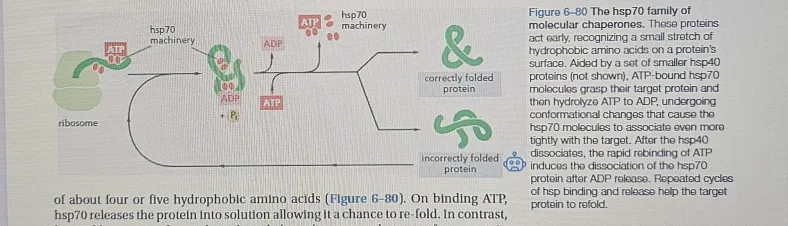
hsp70: Binds to and helps refold misfolded proteins, utilizing ATP.
Early-stage chaperone → assists during nascent protein synthesis.
Prevents aggregation by binding, high affinity, to exposed hydrophobic regions on incompletely folded proteins, and they
hydrolyze ATP, often binding and releasing their protein substrate with each cycle of ATP hydrolysis.
Works co-translationally, meaning it interacts with proteins as they are being synthesized.
Requires ATP for its function:
ATP-bound state → low affinity for substrate (protein).
ATP-bound → Light grip, coaching mode 🏋
ADP-bound state → high affinity, holds onto protein tightly.
ADP-bound → Strong grip, forcing correct form
Repeats the cycle until the protein gets it right ✅
ATP hydrolysis regulates binding/release cycles.
uses ATP to assume different conformations
force incorrectly folded proteins to unfold, gives it another chance to fold correctly.
allows them to attempt proper re-folding
Works with co-chaperones (like Hsp40) to enhance function.
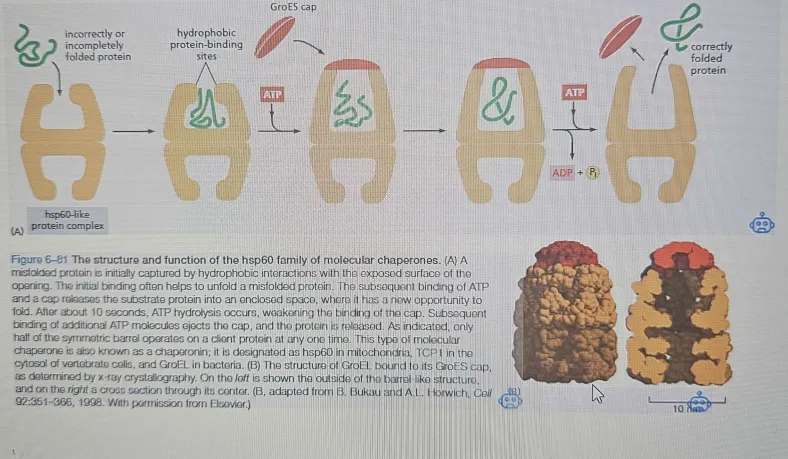
hsp60: Forms a barrel-like structure providing an ideal environment for proper refolding.
Late-stage chaperone → assists in the final maturation of proteins.
Mainly helps signal proteins like kinases and transcription factors.
Functions as a dimer and undergoes ATP-dependent conformational changes.
Plays a key role in stress responses and cancer-related proteins (many mutated oncogenic proteins rely on Hsp90).
provides misfolded proteins an isolated environment in which to attempt proper re-fold
Works in a cycle:
To enter a chamber, a substrate protein is first captured via the hydrophobic entrance. The protein is then released into the interior of the chamber, which is lined with hydrophilic surfaces, and the chamber is sealed
with a lid, a step requiring ATP. Here, the substrate is allowed to fold into its final conformation in isolation, where there are no other proteins with which to aggre-
gate. When ATP is hydrolyzed, the lid pops off, and the substrate protein, whether folded or not, is released from the chamber.
ATP binding → Conformational change → Protein folding → ATP hydrolysis → Protein release.
Quality Control Mechanism
Proteasome-Mediated Protein Degradation: Digest irreparably misfolded proteins.
If refolding fails, proteins are marked for destruction by ubiquitination.
Ubiquitin-proteasome system (UPS) tags proteins with polyubiquitin chains at lysine 48.
Ubiquitin Pathway: Tags misfolded proteins for degradation; large peptides degraded into smaller peptides in the cytosol.
The proteasome is an ATP-dependent multi-subunit protease that degrades proteins into peptides.
Processivity ensures that once a protein enters, it is fully degraded
Proteasome Structure & Function
20S Core Proteasome → Proteolytic chamber with sequestered active sites.
19S Cap → Recognizes and unfolds ubiquitin-tagged proteins before degradation.
AAA+ unfoldase enzymes in the cap thread proteins into the core using ATP hydrolysis.
Protein Turnover and Regulation
Some proteins are permanently short-lived (e.g., damaged/misfolded proteins).
Others are conditionally short-lived (e.g., mitotic cyclins degraded at mitosis).
Regulated protein degradation controls cell cycle progression, stress responses, and metabolism.
Other Infomation
Ribosome as a Ribozyme: Summary for Notes
The ribosome is composed of two-thirds RNA and one-third protein.
rRNA is responsible for the ribosome's structure, positioning tRNAs on mRNA, and catalyzing peptide bond formation.
Ribosomal proteins stabilize the RNA core and aid in initial assembly but don't play a direct role in catalysis.
The 23S rRNA forms the catalytic site for peptide bonds and accelerates the covalent joining of amino acids through precise hydrogen bonding.
The tRNA in the P site contributes an OH group to the catalytic site, ensuring proper positioning for efficient synthesis.
The ribosome is a ribozyme, meaning RNA catalyzes the peptide bond formation, not proteins.
This suggests RNA may have been the first catalyst in early life, and the ribosome's RNA core could be a relic from this time.
Nascent Polypeptide and the Ribosome Tunnel
Ribosome Tunnel:
The nascent polypeptide moves through a water-filled tunnel in the large ribosomal subunit.
Dimensions: Approximately 10 nm × 1.5 nm.
The tunnel walls are primarily composed of 23S rRNA, with a mix of hydrophobic and hydrophilic surfaces.
Function of the Tunnel:
The hydrophobic surfaces provide a "Teflon-like" coating, allowing the polypeptide to slide through without significant interaction.
The tunnel is not complementary to any peptide, allowing easy movement of the polypeptide chain.
Polypeptide Folding:
Nascent proteins are largely unstructured as they pass through the ribosome tunnel.
α-helical regions of the protein may start forming as it exits the ribosome, but full folding into a functional 3D structure occurs later.
Protein Folding:
After exiting the ribosome, the protein must fold properly to achieve its functional conformation.
The process of how proteins fold will be discussed later.
mRNA Quality Control & Surveillance
Eukaryotic mRNA processing occurs in the nucleus before export to the cytoplasm for translation.
Errors in mRNA (damaged, incorrectly spliced, broken) can lead to defective proteins.
To prevent translation of faulty mRNAs:
5’ cap & poly-A tail must be recognized before translation starts.
Nonsense-mediated mRNA decay (NMD) eliminates mRNAs with premature stop codons.
Nonsense-Mediated Decay (NMD) Mechanism
As mRNA exits the nucleus, ribosomes start translating.
Exon junction complexes (EJCs) are displaced as ribosomes move along.
Normally, by the time the stop codon is reached, all EJCs are gone.
If a premature stop codon appears before all EJCs are removed, the mRNA is degraded.
This process prevents toxic truncated proteins.
Role in Disease & Evolution
⅓ of human genetic disorders result from nonsense or frameshift mutations.
NMD helps eliminate faulty mRNAs, reducing disease severity.
May have helped eukaryotic evolution by selecting only functional mRNAs from new gene variants.