Transcription
Learning Objectives:
List and describe the main types of RNA
Describe transcription, including steps in transcription and how the polymerase functions
Define the coding, template, noncoding, nontemplate, and strands
List subunits of prokaryotic RNA polymerase and describe their purposes
List and compare eukaryotic and prokaryotic promoter elements
Describe the function of sigma factor, and how sigma factors are used to control transcription.
Describe RNA proofreading mechanisms
Describe termination in prokaryotes. Compare intrinsic and rho-dependent termination
Describe TFIID and TFIIB in eukaryotes. Explain how they help initiate transcription
Transcription is copying the information in DNA into RNA:
Makes it useful to make proteins
Central dogma
Most RNA is not translated
Based on volume, the vast majority is not.
most not made into new proteins
Three types of RNA:
rRNA: Ribosomal RNA, makes up the RNA, and carries out the chemical reactions in the ribosome
Run the ribosome in electrophoresis, get two big bands, which is rRNA; the rest is very small
mRNA: Messenger RNA, stuff that gets transcribed into RNA, goes to ribosomes and makes the proteins. Messenger between the DNA and the ribosome
Most regulated and the most changing
tRNA: Transfer RNA, the smartest molecule; speaks both nucleic acid and amino acid
An adaptor molecule in between mRNA and the protein
The most prominent way of regulating a particular gene pathway is through transcription
regulate whether it gets transcribed and still even then weather it gets translated, and if protein is made, whether or not it works
Want gene, turn on RNA pol, to make the gene
Based on energy, and regulates on the transcriptional level to save energy on the other levels
Clotting factors different, becuase we could be dead without it, so it gets cleaved.
Energy efficent or fast
All comes back to DNA
RNA:
Is a polymer, like DNA
Different from DNA because it has the extra OH group on the 2’ carbon
Single-stranded, doesn’t have a backup copy of itself
very short-lived and can just make another copy of itself
it can fold itself up, base pairs within the molecule
more reactive than and more unstable than DNA, so can do more chemical reactions
Different RNA’s have different purposes
Think that there’s mRNA, tRNA, and rRNA all with different jobs, DNA has the same job across the board
THINK: Active, transient, reactive, and alays doing stuff
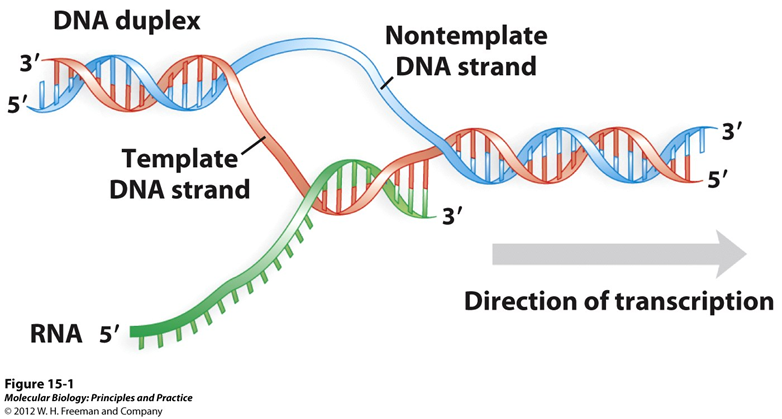
Strand names:
Each strand being copied and strand not being copied has a name and a nonname
Sequence of RNA is going to be the same as the coding strand
ALWAYS 5’-3’
Code in DNA is the same as in the RNA
Also, the nontemplate strand becuase it’s not the templat
Template strand: what is actually physically being coipied by the RNA
Also, known as the noncoding strand
RNA is made by RNA polymerase:
RNA polymerase basically does the same thing as DNA polymerase
The chemical reaction that forms DNA is the same that forms RNA
Attacks 3’ hydroxyl group
RNA uses uracil instead of thymine
everywhere where there is a T in DNA there’s a U in RNA
Because RNA is constantly being made and destroyed if the C becomes a U, if it happens a couple of times it’s not a big of a deal
RNA polymerase doesn’t need a primer
Prokaryotic RNA polymerase:
β, β’ subunits: where the RNA catalysis happens, where nucleotides come together
ω subunit helps with assembly
α subunits: Help with assembly and also regulatory, bind to other proteins
regulatory and structural
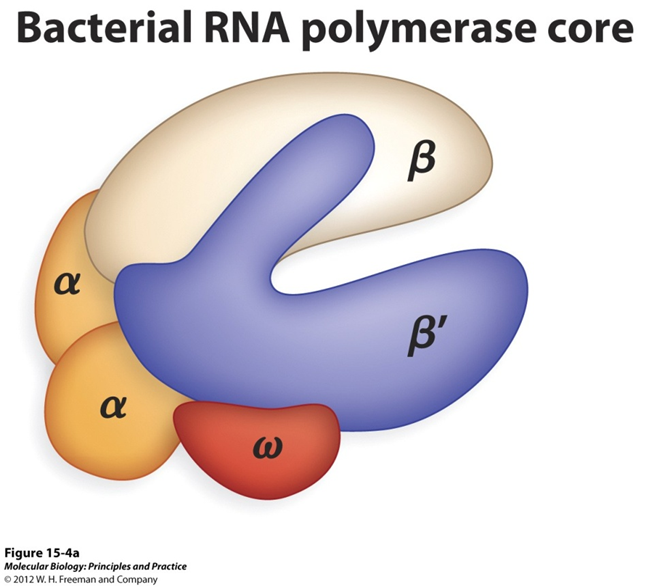
Eukaryotic RNA polyemerase is similar to prokaryotic
is bigger
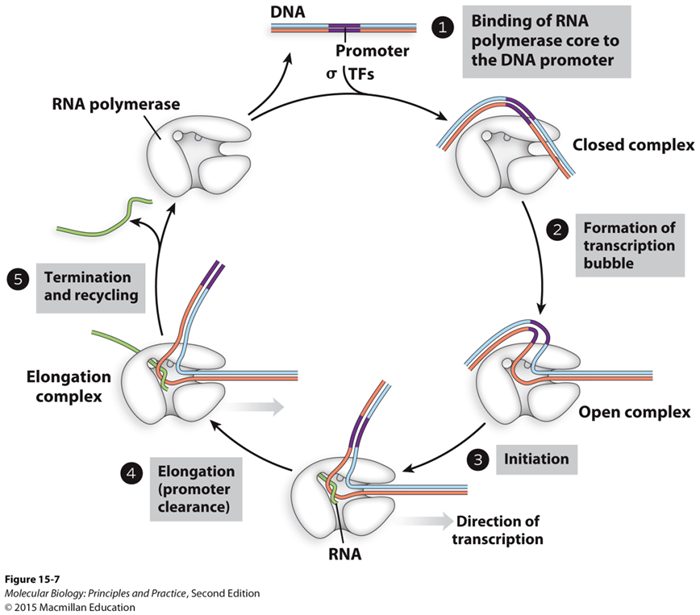
Cycle of transcription:
Initiation:
The start of transcription
Starts at the closed complex
Have a promoter and RNA polymerase stuck together
Promotor: sequence of DNA that RNA polymerase recognizes
Then become an open complex
RNA polymerase goanna take that DNA
and open it up a little bit
forming the transcription bubble
Promoter:
DNA sequence that is recognized by RNA pol. 2
Mark’s beginning of transcription
NOT LIKE A START CODON
Can be different promoters
Could have a strong promoter for genes you want on all the time
could have weak promoters that aren’t bound to the polymerase often for genes you don’t want on all the time
So less RNA is being made
A better promoter is recognized by the RNA, the more transcription that can occur
Position and orientation dependent
The way the RNA polymerase faces determines which direction the RNA polymerase goes
Sometimes, different combinations of DNA sequences that make up the promoter
Bacterial promoter:
The core promoter has a -10 box and a -35 box
That many nucleotides upstream from where transcription starts
bacterial promoter is 10 and 35 nucleotides upstream from the start of RNA transcription
25 nucleotides apart because that’s the amount of DNA it takes to go all the way around
+1 site where transcription starts
goes from -1 to +1 where transcription starts
UP element: not necessary for transcription, but an A-T-rich sequence of DNA that helps transcription begin,
Helps RNA pol. to bind the promotors
Have a -35 box, a -10 box, and an UP element will make a pretty strong promotor, and will get lots of RNA
Consensus Sequences:
If you took all the promoters in all bacterial sequences and went, what’s the most common nucleotide at -10, -11, or -13, that’s the consensus sequences
Standard sequences for the promoters
-10: TATAAT
-35: TTGACA
If the standard sequence is then, RNA polymerase will be able to recognize, and it will be a strong promoter
So closer to the exact sequence, the better it will be at transcription
Somehow modify something to cause transcription to stop or start
Most promoters don’t function unless you do something to make it work.
Sigma factor:
Is the intermediary factor that the RNA polymerase binds to
Recognizes and binds to the promoter
Sigma 70 default factor
If in another state, it can change to where it would recognize a different -10 and -30 promotor box, to where it can become resistant to whatever is trying to kill it
Need both regular and heat-shock promoters, then you have two promoters, one for each
Use if you want to change your entire transctopional system
Go from one set of genes to another to get another sigma factor
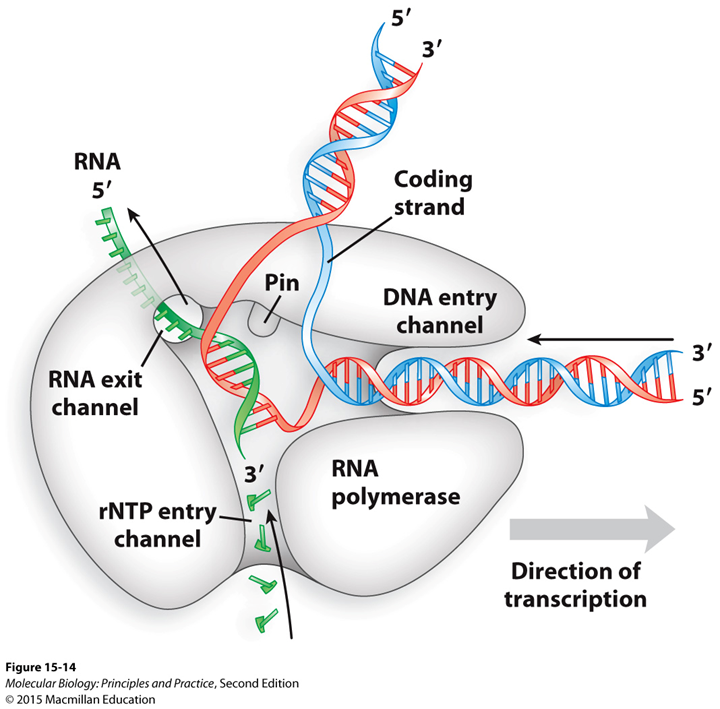
Structure-function of RNA polymerase (elongation):
Making the RNA
Have an entry channel where DNA polymerase takes in DNA, or RNA pol. is moving along DNA in that direction
Pin pulls DNA apart and holds apart
RNA exist channel is where RNA that is being synthesized exits
The rNTP entry channel is where the nucleotides enter in.
Once made 6-8 nucleotides will hold RNA pol on DNA, so it can keep going
Kinetic Proofreading:
Depends on speed/rate of reaction
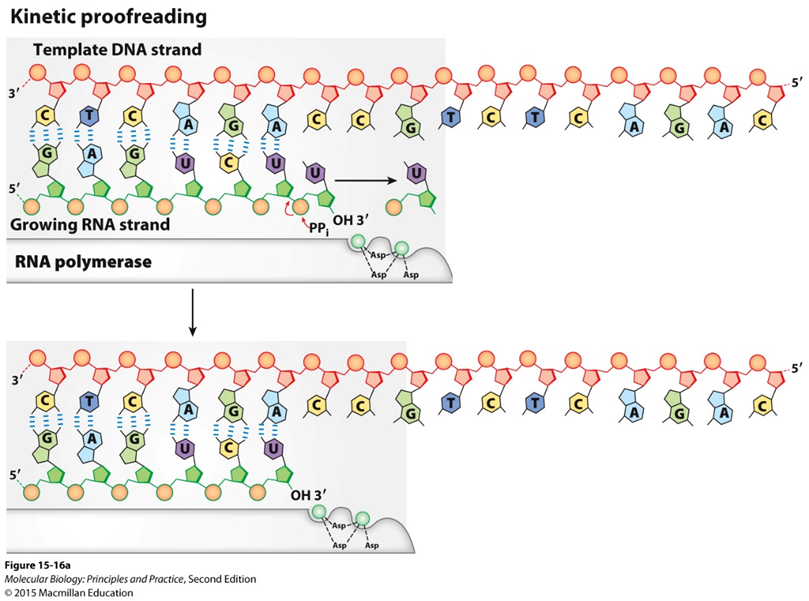
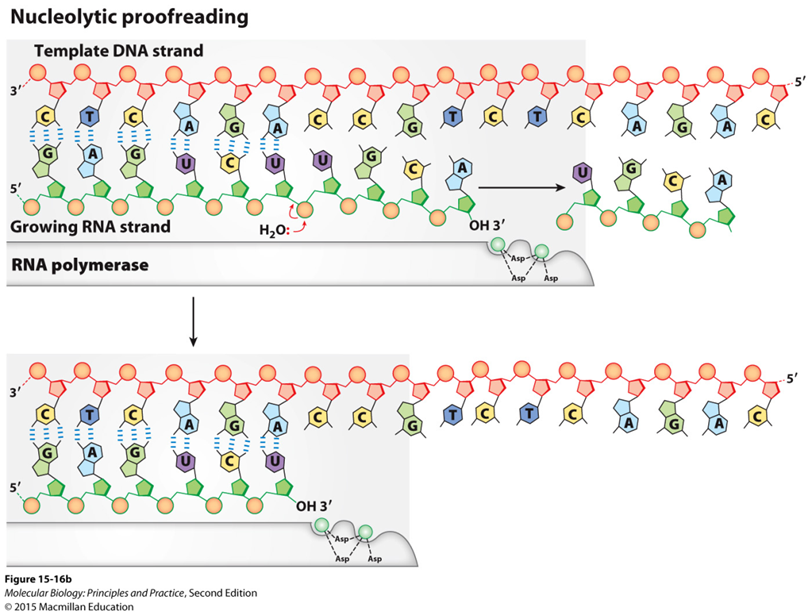
Nucleolytic proofreading:
RNA polymerase is zipping along when all of a sudden it makes a mistake, and has a internal nuclease
cut out a whole chunk of nucleotides that were made and repairs the mispair
Lysing, cutting the chain
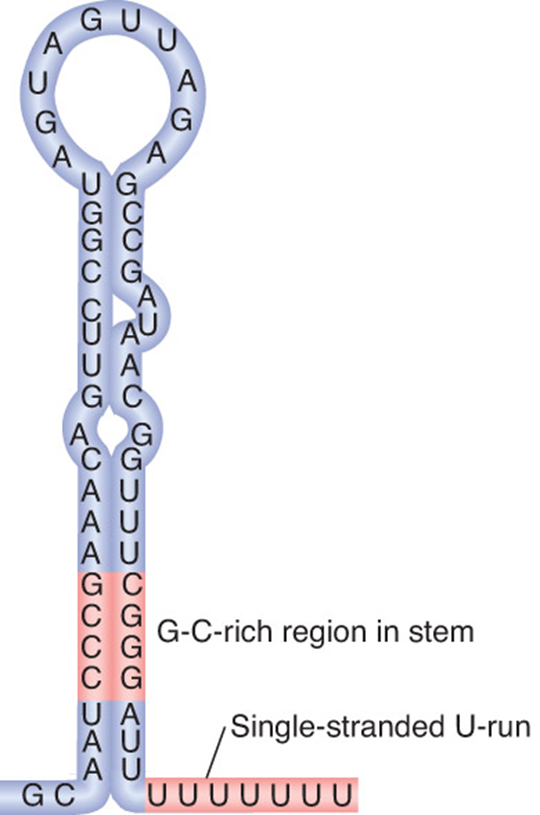
Termination:
End of transcription
RNA pol. needs to know when/where to stop
Need to have a stop signal
2 ways:
intrinsic termination
hairpin: folds up on itself
and Rho-dependent termination
Protein inside bacterial cells
Jumps on when there is a rut
races after RNA Pol. and then once it reaches a hairpin, Rho is able to catch up, and it will kick RNA pol. off the DNA
Both use a hairpin structure
Eukaryotic Transcription:
Three different polymerases:
Polymerase 1: rRNA
Mostly makes rRNA, very busy
Polymerase 2: mRNA
transcribes messenger RNA, most regulated out of the bunch
Polymerase 3: tRNA
transcribes tRNA, decodes mRNA
All are regulated through:
Basal and specific transcription factors:
Transcription factor: a protein that binds to DNA and affects transcription
Different for each gene, different combinations
Basal transcription factor: required for all transcription
EX: TF2D
promoters are specific fo each polymerase
Eukaryotic Promoters (pol II):
Where RNA pol. binds and starts in prokaryotes
4 possible promoter elements in Eukaryotes:
The TATA box
B response element (BRE
Downstream promoter elements
or the initiator (INR)
Have to have at least 2 of the four for it to successfully start transcription
Have to have the initiator and the TATA box for it to work
Mostly get bound to general transcription factors in prokaryotes
Eukaryotic basal transcription factors (pol II):
TF2D is a complex of differnt proteins so it can bind to different things such as the TATA box or the initator
So why need two to allow the factor to bind and do its thing
Once binds recruits TF2B which then binds to TBP, and positions the polymerase