Restriction Enzymes
Definition of Restriction Enzymes
Restriction enzymes, also known as restriction endonucleases, are a class of enzymes that cleave DNA in response to specific sequences known as recognition sites located within double-stranded DNA ( recognition site ). This property allows them to cut DNA at precise locations, facilitating various molecular biology applications.
Types of Restriction Enzymes
Restriction enzymes are essential components of bacterial defence mechanisms against foreign DNA, such as bacteriophage DNA. They can be categorized into three main classes:
Types I & III: These are restriction-modification enzymes that cut DNA either upstream or downstream of their recognition sites.
Type II: This class is the most significant for molecular biology applications, as they cut DNA directly within their recognition sites or very close to them.
Action for Type II Restriction Enzymes
Approximately 3000 bacterial restriction enzymes have been identified across varied species. The primary feature that distinguishes each enzyme is its recognition site.
The recognition site is a short sequence, typically containing 4 to 8 base pairs (bp), within the total DNA sequence, most often configured as an inverted palindrome.
Example of a Recognition Site: The recognition sequence for EcoRI is:
Forward: GAA TTC
Reverse: CCT TAG
Methylase Attendant to Type II Enzymes
Every bacterial species that possesses a restriction enzyme also possesses a corresponding methylase enzyme.
This methylase modifies the sites in its genome to prevent its own DNA from being cut by the restriction enzyme.
For instance, E. coli produces the EcoRI restriction enzyme along with EcoRI methylase.
Example with EcoRI
The cleavage by EcoRI occurs at the recognition site's central nucleotide pairs:
Cleavage sequence: 5'-GAATTC-3' results in sticky ends:
5' sticky end: GAATTC
3' sticky end: CTTAAG
The methylated form of the DNA is not cleaved by EcoRI, as indicated by the methylation marking (m) on the DNA sequence.
Characteristics of Different Restriction Enzymes
A comparison of certain restriction enzymes based on their recognition sites, cleavage positions, and the type of overhanging ends they produce includes:
HinPI
Length: 4 bp
Recognition Site: 5'-GCGC-3'
Type of Ends: 3'-overhang
EcoRI
Length: 6 bp
Recognition Site: 5'-GAATTC-3'
Type of Ends: 5'-overhanging
PstI
Length: 6 bp
Recognition Site: 5'-CTGCAG-3'
Type of Ends: 5'-overhanging
Sspl & NotI
Length: 8 bp
Recognition Site: Specifically cleave and generate flush or blunt ends respectively.
Nomenclature of Restriction Enzymes
The naming convention for restriction enzymes is derived from their bacterial origin and the order of their discovery. For example, the enzyme HaeIII is named from:
First initial of genus (Haemophilus)
Roman numerals indicating the discovery order
First two initials of species (aegypticus).
Strain Designation in Enzyme Nomenclature
If an enzyme is sourced from a specific strain of bacteria, the three-letter species designation may be followed by a strain identifier.
Calculating Restriction Site Frequency
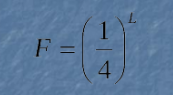
L: Length of the recognition site
F: Frequency of occurrence of the recognition site
The theoretical frequency calculation assumes a random incorporation of nucleotides during DNA synthesis. However, in nature, certain nucleotide compositions may be selected for specific functions, leading to unequal distributions. For instance, genomic regions rich in expressed genes often exhibit a higher G/C content in many higher eukaryotes.
Practical Implications of Restriction Site Frequency
An analysis comparing estimated site frequencies (F) and actual frequencies in bacteriophage lambda DNA (45,000 base pairs) provides insight into the distribution of restriction sites.
Sticky Ends and Cohesive Termination
When restriction enzymes cleave DNA, they can leave either 5' or 3' overhangs, commonly termed "sticky" or cohesive ends. These ends can re-anneal with any complementary DNA sequence due to base-pairing, provided the conditions favor re-annealing. This property allows for varied applications, including the formation of recombinant genes, adding adapters to DNA, or cloning DNA fragments into vectors.
Applications of "Sticky" Ends
Two DNA fragments with compatible ends can temporarily pair by hydrogen bonding; for permanent linking, a DNA repair enzyme is required.
Applications include creating recombinant DNA molecules, ligating ends to generate circular DNA from linear fragments, and more.
Inverse PCR
Inverse PCR is an approach used when a known DNA sequence is surrounded by unknown regions, allowing for amplification moving outwards from a known region by designing primers accordingly. Post cutting the DNA with restriction enzymes, ligation creates a circular DNA molecule, facilitating inward amplifications during PCR.
Isoschizomers
Isoschizomers are different restriction enzymes that recognize the same DNA sequence. There are three categories:
Perfect Isoschizomers: Recognize and cleave the same sequence (e.g., NsiI and AvaIII).
Imperfect Isoschizomers: Share recognition sites but cleave at varied points, resulting in differing sticky ends.
Methylation-sensitive Isoschizomers: Differ in cutting ability depending on whether bases are methylated, useful for assessing methylation patterns in DNA.
Generating Compatible Ends
Different restriction enzymes may act on distinct cleavage sites, producing compatible cohesive termini. This allows for hybrid site formation by ligation of overhangs generated from different enzymes. Example: Population of two cut enzymes resulting in a hybrid sequence:
PstI and NsiI joined could yield a combined sequence: 5'-CTGCAT-3' and 3'-GACGTA-5'.
Incorporating Restriction Sites into PCR Primers
To ensure a DNA segment has designated restriction sites, they can be included within the PCR primers used for amplification, provided that these sites do not appear elsewhere within the segment.
Assessing Genetic Diversity Using Restriction Enzymes
Recognition sites being specific means any mutation altering a site will prevent digestion by the enzyme, thus resulting in varied restriction profiles across different individuals. This variability can be evaluated through techniques like:
RFLP (Restriction Fragment Length Polymorphism)
AFLP (Amplification Fragment Length Polymorphism)
RAD (Restriction site Associated DNA)
AFLPs - Amplification Fragment Length Polymorphisms
These techniques utilize varying primer combinations to yield distinct DNA fragment profiles, crucial for assessing genetic diversity across populations.
Analysing Restriction Fragments
Assessment of restriction patterns can be done through various fragments generated, depending on whether no digestion, partial digestion, or complete digestion is undertaken.
Example of PCR-RFLP: In the example involving the Lipase Maturation Factor 1 (LMF1) gene mutation, specific mutations result in different numbers of bands when digested with AvaI:
If the sequence is T, a single band is produced.
If C, two bands are produced.
Mixed alleles lead to three bands total.
Generating a Simple Restriction Map
An analysis of the λ DNA (48.5 Kb) reveals clear distinctions between single and double digestions, which yield fragments of defined sizes. Sizes of fragments can be both essential in understanding the structure of the DNA and determining the presence of specific recognition sites across different DNA samples.
Single and Double Digestion Fragment Details
Fragments resulting from various restriction enzymes can be laid out in terms of size and count, providing insights into the DNA structure.
Conclusion
The knowledge and understanding of restriction enzymes pave the way for numerous applications in research, including cloning, genetic engineering, and diagnostic techniques widely utilized in molecular biology and genetics.