Lecture 27- Diagnostic Testing
Diagnostic Testing for Disease Variants
Learning Objectives
Explain how PCR and PCR-RFLP can be used in genetic disease diagnosis.
Interpret results of PCR and PCR-RFLP diagnostic tests.
Design a protocol for a diagnostic test, including control experiments and examples.
Describe some examples of diagnostic testing applications.
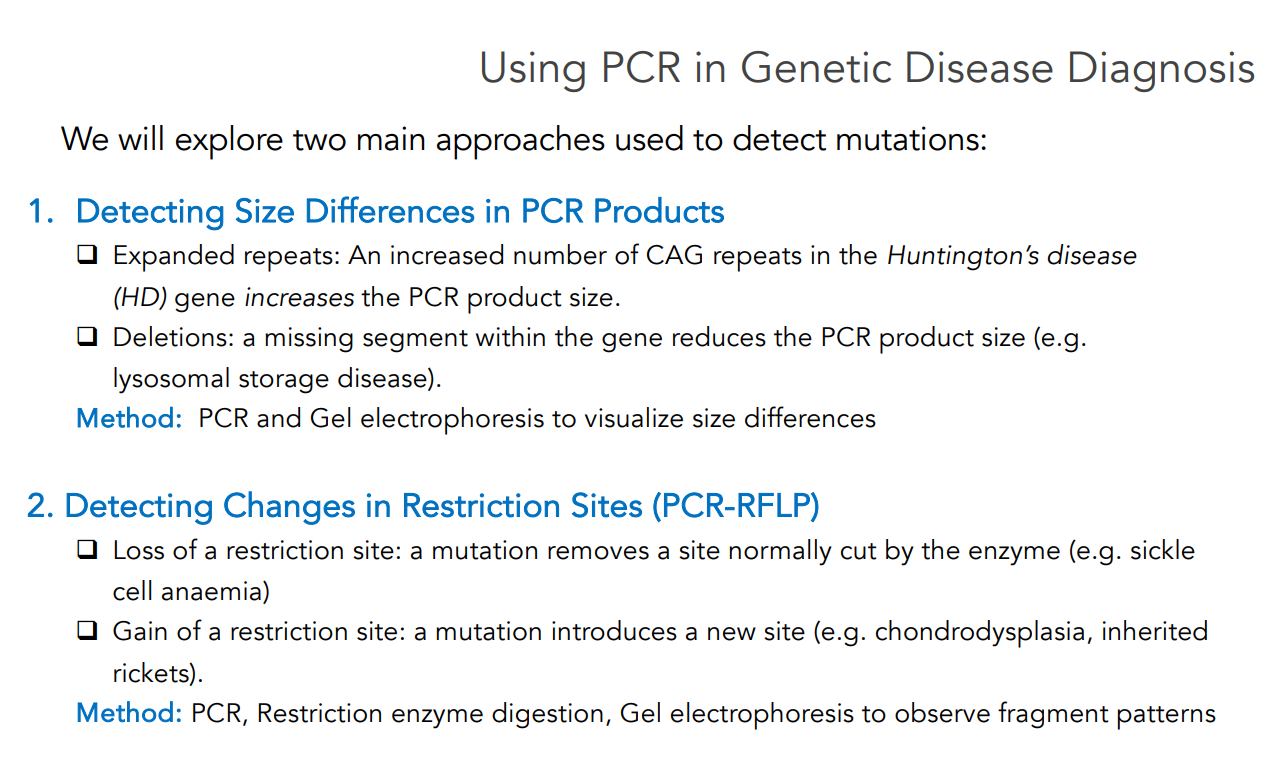
Using PCR in Genetic Disease Diagnosis
Two main approaches that are used to detect mutations:
Detecting Size Differences in PCR Products
Certain mutations change the size of the DNA fragment amplified by PCR.
Expanded repeats: An increased number of CAG repeats in the Huntington’s disease (HD) gene increases the PCR product size.
Deletions: A missing segment within the gene reduces the PCR product size (e.g. lysosomal storage disease).
Method: PCR and Gel electrophoresis to visualize size differences
Detecting Changes in Restriction Sites (PCR-RFLP)
Some mutations affect recognition sites for restriction enzymes.
Loss of a restriction site: A mutation removes a site normally cut by the enzyme (e.g. sickle cell anaemia)
Gain of a restriction site: A mutation introduces a new site (e.g. chondrodysplasia).
Method: PCR, Restriction enzyme digestion, Gel electrophoresis (to observe fragment patterns)

Why Genetic Testing?

Why PCR-Based Tests?

Accommodates small amounts of DNA.
Rapid testing.
Fast transitions from the lab to the clinic.
Diagnostics is now an > billion global market.
Diagnosing DNA Size Changes
DNA from a single individual is run in each lane.
Stained ‘bands’ indicate PCR products with different sizes.
Different sized PCR products represent different genetic alleles.
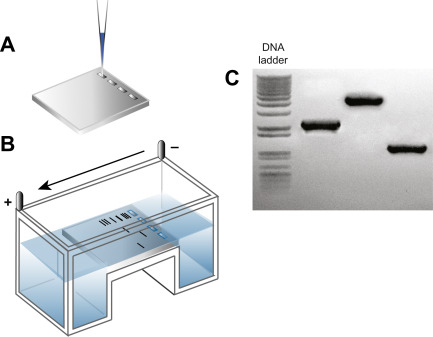
Interpreting Results: Detecting Size Differences in PCR Products Example of Expanded Repeats: Huntington’s Disease (HD)
Huntington’s disease
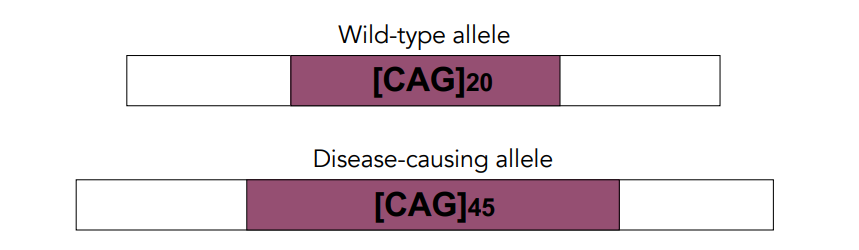
A dominantly-inherited disorder caused by extra copies of a glutamine codon (CAG) in the Huntington’s disease gene (exon 1).
People with 40 or more CAG repeats will develop the disease.
A larger number of repeats is associated with an earlier age of disease onset.
PCR can be used to distinguish these alleles based on length.
Large band (disease).
Small band (wild-type).
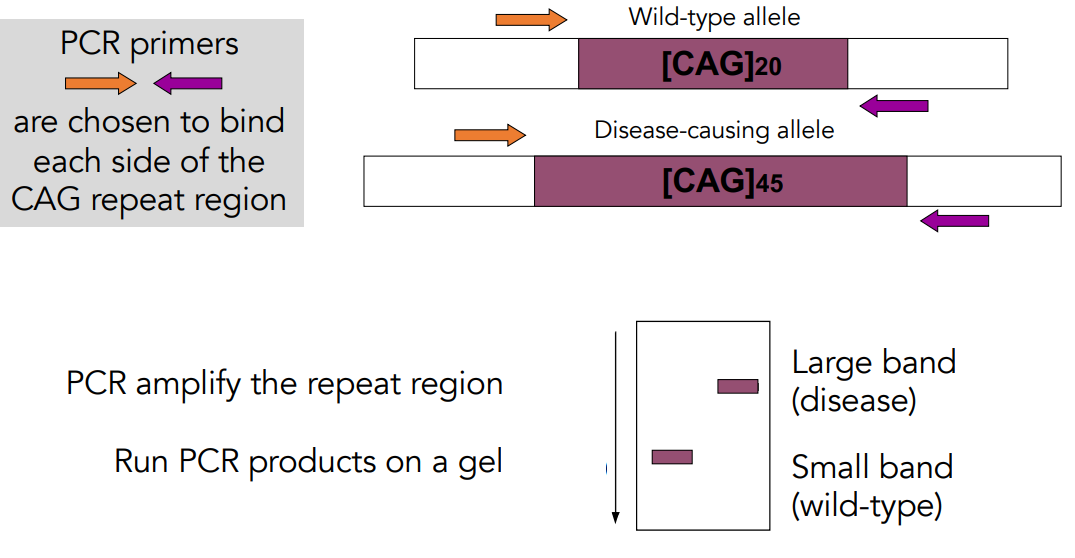
A larger number of CAG repeats is associated with an earlier age of disease onset.
The CAG repeat mutation is unstable; repeat expansion can occur with parental(from the father) transmission.
A smaller number of CAG repeats is associated with later age of onset.
Interpreting Results: Detecting Size Differences in PCR Products Example of Deletions: Lysosomal Storage Disorder
Lysosomal Storage Disorder
A metabolic disease leading to a build-up of toxic materials in cells.
Lesions occur in many cell types, including corneas (eyes), as complex sulphated sugars are not broken down in lysosomes.
A 22 bp deletion in arylsulphatase B gene caused a frame shift and premature stop codon.
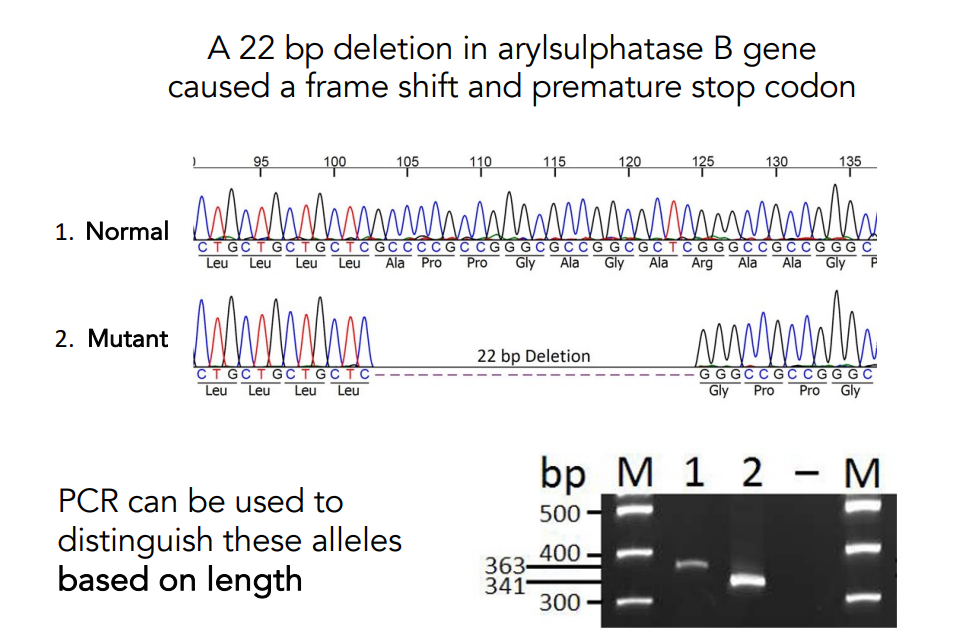
PCR can be used to distinguish these alleles based on length.
Interpreting Results: Detecting Changes in Restriction Sites (PCR-RFLP) Example of a Loss of a restriction site: Sickle Cell Anemia
Sickle Cell Anemia
A severe hereditary form of anemia.
A mutated hemoglobin distorts red blood cells into a crescent shape at low oxygen levels.
Common among individuals with African ancestry.
Wild Type vs. Sickle Cell
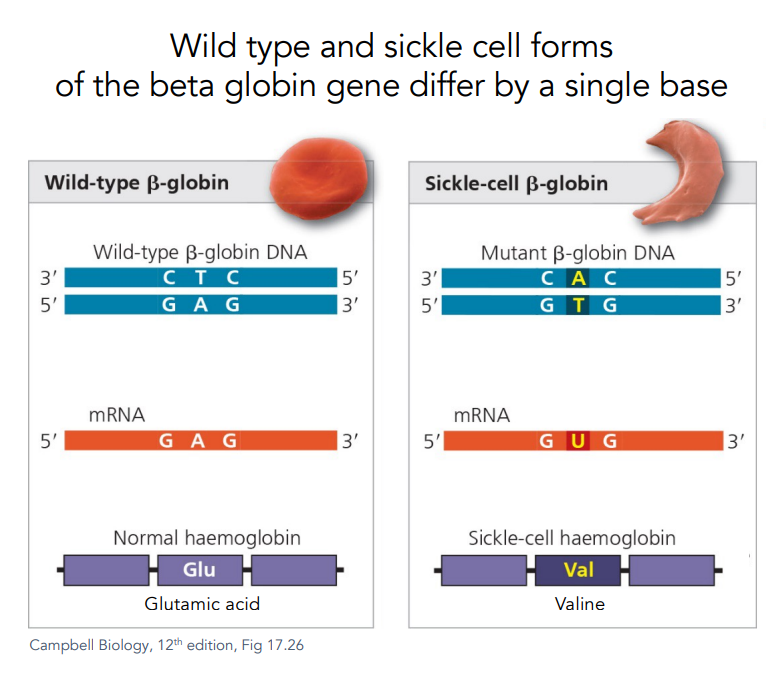
Wild type and sickle cell forms of the beta globin gene differ by a single base.
Glutamic acid is replaced by Valine.
Procedure Order for Sickle Cell Anemia Test
Correct order of procedures:
PCR amplify the DNA
Cut DNA with DdeI(Restriction Endonuclease)
Run DNA on gel
Restriction Endonucleases (Ddel)
An enzyme that cuts DNA at specific sequences
Bacterial defense system.
Cut at a specific symmetric sequence (e.g., EcoRI cuts at GAATTC).
Degrade invading DNA (e.g., from phages).
Does not cut the bacteria’s own chromosome (bases are methylated).
Mutation and Restriction Enzyme Sites
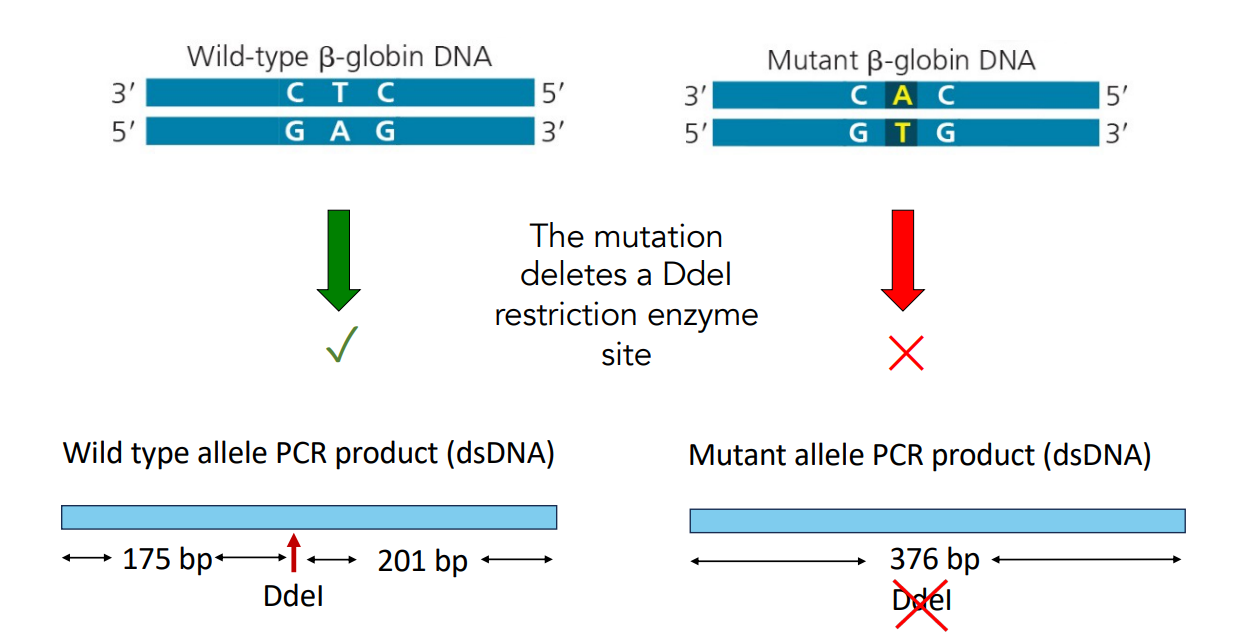
The mutation deletes a DdeI restriction enzyme site.
Wild type allele PCR product (dsDNA) is 201 bp.
Mutant allele PCR product (dsDNA) is 376 bp.
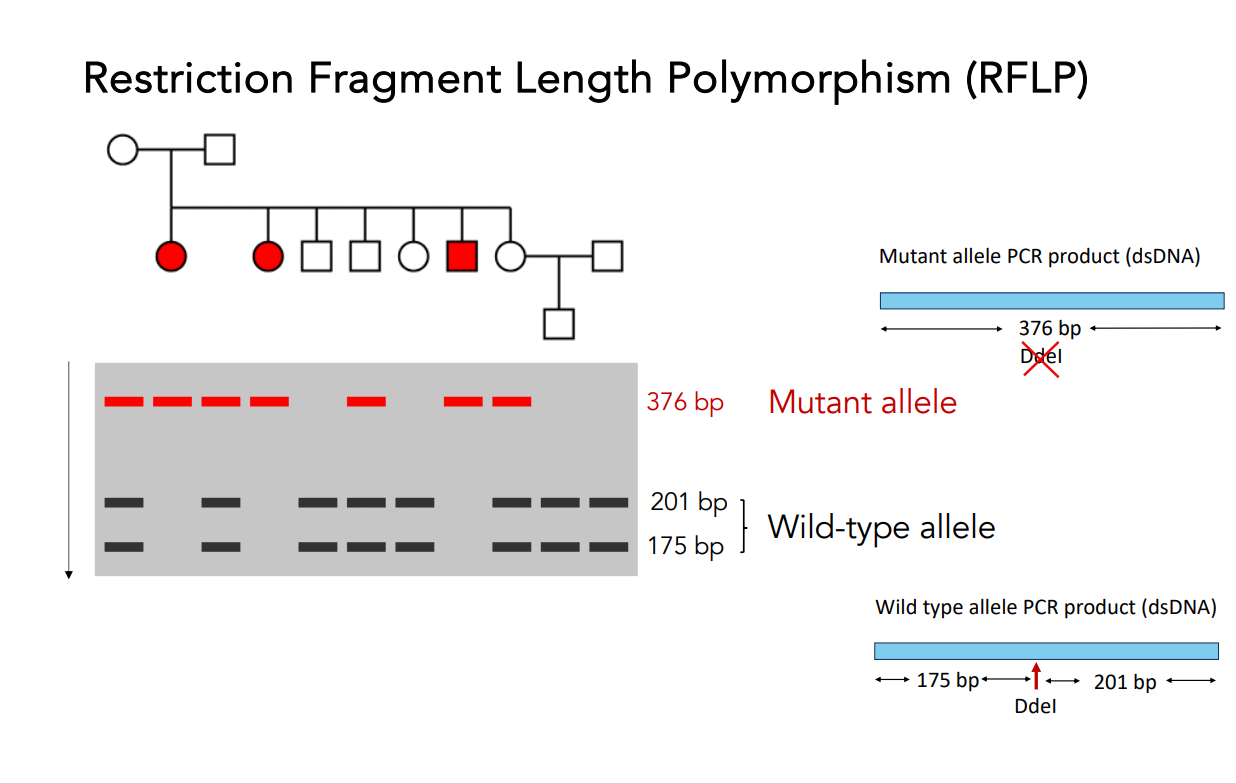
Interpreting results
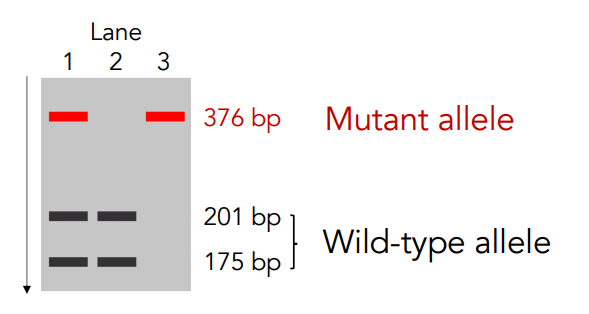
Interpreting Results: Detecting Changes in Restriction Sites (PCR-RFLP) Example of a Gain of a restriction site: Chondrodysplasia
Chondrodysplasia
Malformation of the cartilage.
Impairs normal development of many parts of the body.
Dwarfism, barrel-shaped chest, wide-based stance, shortened neck, forelimb deformities, deformed cartilage of the trachea

Genetic Cause of Chondrodysplasia in Texel Sheep
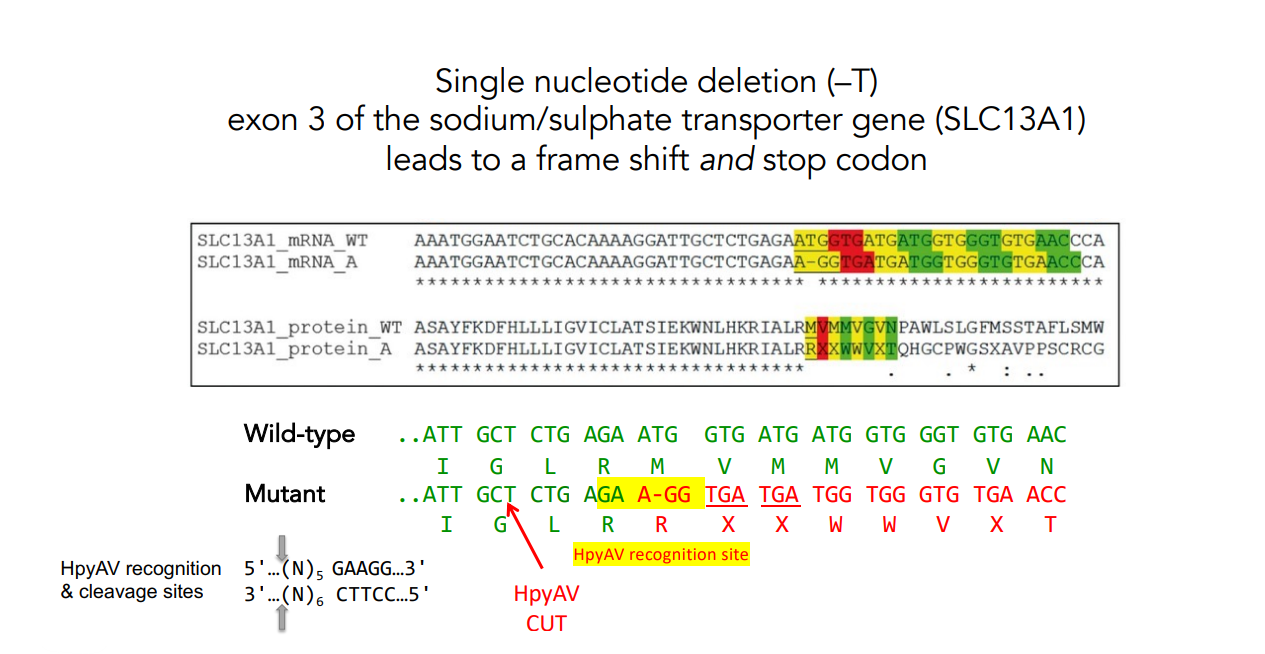
HpyAV Recognition Site and PCR-RFLP for Chondrodysplasia
The single base pair deletion (–T) creates a new recognition site for restriction enzyme HpyAV.
PCR-RFLP can distinguish these alleles (i.e., PCR the DNA, then cut with HpyAV).
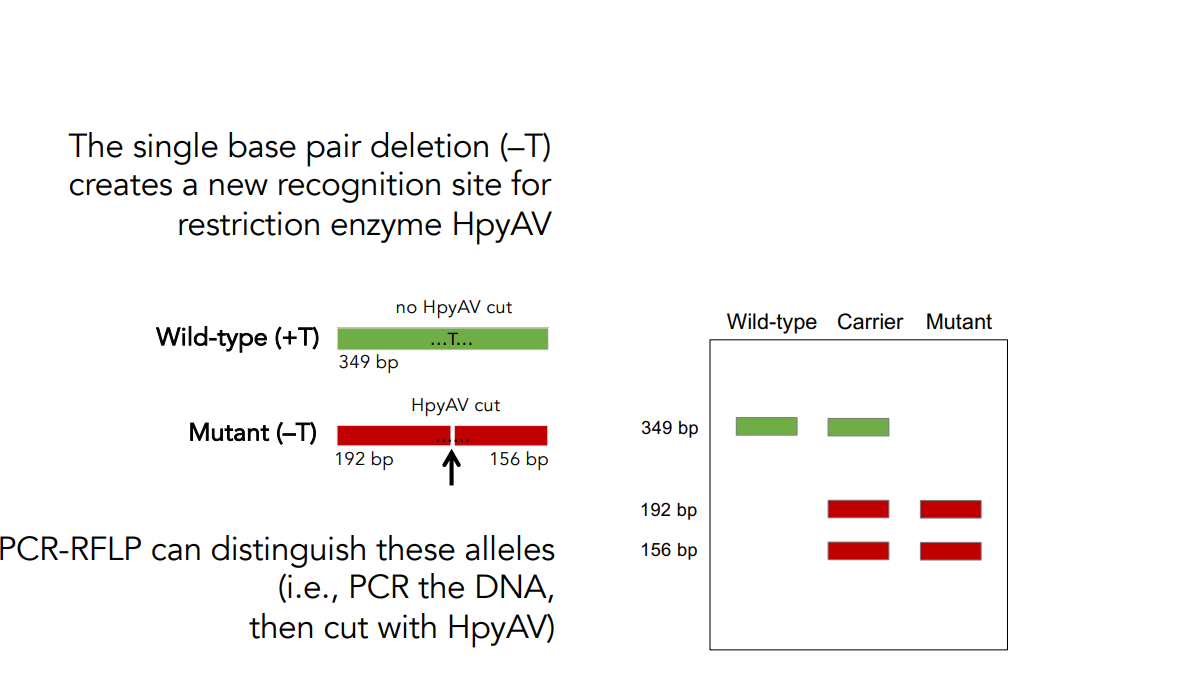
Wild-type (+T) has no HpyAV cut.
Mutant (–T) has HpyAV cut.
Draw the expected pattern of DNA fragments on a gel after PCR-RFLP.