EXAM2 GENESANDGENOMES
Endicott College
Genes and Genomes
Unit II Study Guide
Molecular Genetics
Know what a DNA secondary structure is and its defining characteristics.
Be able to give an example
Double-helix structure; two complementary strands forming a right-handed helix
Each strand had nucelotude sequences
Antiparallel 5 - 3 to 3 - 5
Base pairing via hydrogen bonds
Linking of the nucleotides down the strand
Phosphodiester bonds; 5’ phosphate group to the 3’hydroxyl group of the next nucleotide
The backbone of DNA and RNA is hydrophilic
Hairpin and Cruciforms
Repetitive sequences bending DNA
What base pairing is harder to break apart? AT or GC?
GC because of hydrogen bonding. 3 hydrogen bonds in GC vs. 2 hydrogen bonds in AT
Know the concept of denaturation and applications of its use in molecular biology
Since DNA is highly viscous (or sticky and glue-like) with a pH of 7.0, when denatured (hydrogen bonds are broken down, helix unwinds, and DNA strands are separated) this is done by HEATING the DNA to make it less viscous
DENATURATION
DNA slowly denatures when heated at a slow rate
Double strand — break (melting point) — single strand
Can be renatured aka ANNEALED
PROBING WITH DNA AFTER DENATURATION
Once denatured it can be glued together aka annealed with exogenous or outside DNA
Probes – short DNA sequences that is complementary to a gene of interest
Know what a Southern blot measures
Southern blot detects specific DNA fragments
Know how restriction enzymes work
Restriction enzymes recognize specific DNA sequences
They are nucleases that cleave DNA between nucleotides
They’re like molecular scissors because recogzine unique short sequences of DNA and cut the DNA at those exact locations
Recognition: Each restriction enzyme has a unique DNA sequence it recognizes. For example, the enzyme EcoRI recognizes the sequence GAATTC.
Cutting: When the enzyme finds its specific sequence on a piece of DNA, it cuts the DNA at or near that sequence. This cut can create "sticky ends" (overhanging single-stranded DNA) or "blunt ends" (straight cuts with no overhang).
Results: Once the DNA is cut, the fragments can be used in different ways. For example, sticky ends allow the DNA to be easily joined with other DNA pieces cut by the same enzyme, which is useful for creating recombinant DNA.
Be able to identify the steps of PCR and understand the exponential amplification principle.
DENATURATION
Heating
Break hydrogen bonds
Separate DNA strands from double to single-strand
ANNEALING
Cooling
Adding in short DNA primers or exogenous DNA to complementary sequences on the single-strand
EXTENSION
Temp is raised
Heat stable DNA polymerase, Taq polymerase, synthezies a new DNA strand
Starts at primers then works its way down in the 5’ to 3’ direction
Complementary strand is made to each template strand
The principle of exponential amplification in PCR is based on the fact that with each cycle, the amount of DNA is approximately doubled.
In the first cycle, one DNA molecule produces two copies.
In the second cycle, those two copies are each duplicated, resulting in four copies.
By the third cycle, you get eight copies, and so on.
N = N0 x 2^n
HOLD
Know what qPCR is.
REAL-TIME PCR
Specialized version of the PCR that enables the quantification of DNA in real-time
Traditional PCR tells you if DNA is present after ALL cycles are done
Type of big question I might ask:
None, this lecture will be incorporated into other
questions. I might give you a picture of a gel to interpret or give you an experimental
situation where I give you choices and you must pick the right method and explain why.
DNA and RNA are nucleic acids that carry our genetic information
Nucleic acid structure is tied to the chemistry of each nucleotide
DNA is a right-handed B-form double helix but can form alternative DNA structures and helices
RNA is not always a linear molecule and relies on base pairing
Classic molecular biology techniques rely on key features of the structure of nucleic acids and have revolutionized science
Di-deoxynucleotide
First-generation (Sangar) DNA sequencing which relies on di-deoxynucleotide terminators
Has Hydrogen on its 3’ carbon (or hydroxyl group) which stops the chain from elongating aka stops sequencing
Sanger Sequencing and next-generation sequencing
Know how Sanger sequencing works and be able to read a traditional Sanger sequencing gel.
Chain termination (Sangar) sequencing
“Endpoint PCR”
4 large reactions
Large gel
Radioactivity
Polyacrylamide gel electrophoresis
PCR with fluorescent chain-terminating ddNTPs
Size separation by polyacrylamide gel electrophoresis
Laser excitation and detection by sequencing machine
Why it’s better than traditional gel electrophoresis
More sensitive
More quantitative
Faster
Be able to tell the difference between a regular nucleotide and a di-deoxynucleotide.
Regular Nucelotide
OH on the 3’ carbon, continues the chain and sequencing
Di-deoxynucleotide
H of the 3’ carbon, ends the chain and stops sequencing
Know the applications of Sanger vs. next generation sequencing( Illumina/Ion Torrent/ Pyrosequencing) (what would you use if given a scenario)
Chain terminating Sangar/qPCR
Single site and single gene
Next generation sequencing (NGS)
Single site and single gene
Multi-gene to whole genome
Know the steps of library prep
Input DNA
Fragmentation
Adaptor ligation
Sequencing
Know the major differences between the Next Gen Methods
Illumina:
Flourscenence of 1k bases added simultaneously to a cluster of blonded sequences
Ion Torrent
pH proportional to H+ concentration
Pyrosequencing
Chemiluminescence proportional to pyrophosphate concentration
PacBio
Flourescence of one base (ZMW)
Be aware of the major bioinformatic steps of WGS. (NGS??) 3 major phases SAF
Sequencing
Get sequence and stitch together contigs
Assembly
Sequence more and get longer contigs until the sequences are ordered for each chromosome
Maybe some unknown sequences
Finishing
You have the whole sequence/ genome tbh
Only fruit fly, human, and mouse genomes are at the finish phase
Know the uses of NGS—what have we been able to use it for?
Major point: sequence a lot of DNA in a short period of time
Genomes from individual cancer cells
Complex mixtures of cells
Determine the nature of mutation and evolution
PGD, disease discovery, GWAS
GWAS – decided to genotype 12 million common SNPs
Genotyped all DNAs for all SNPs
Trying to investigate the heritability of common diseases
What is Telomere to Telomere assembly? What regions of the genome have been a gap in the human genome?
GWAS
What is a GWAS? What is the goal of a GWAS?
GENOME WIDE ASSOCIATION STUDIES
GOAL:
TEST WHETHER GENETIC POLYMORPHISMS (ALLELES) ARE ASSOCIATED WITH DISEASE STATUS
AND
FIND GENETIC MARKERS AT WHICH VARIATION IN GENOTYPE IS SIGNIFICANTLY ASSOCIATED WITH VARIATION IN PHENOTYPE OF POPULATIONS
Why has GWAS gained popularity since the early 2000s? What type of Biology research does it fall under?
CASE-STUDY CONTROLLED STUDY DESIGNS and falls under systems biology AND functional genomics but it comes at it with a REVERSE GENETICS APPROACH
It is popular because we are gaining knowledge about the context of human disease
Computer science has been on the rise as well as bioinformatics, the collection of large data sets, and the rise of genome sequencing
What are SNPs
Single Nucleotide Polymorphisms
Genetic differences between INDIVIDUALS
IMPACT
No impact
Health issues
Most common type of genetic variation
Polymorphism
Variations in the DNA sequence that are common in a population
Not always linked to negative outcomes like mutations
Represents genetic diversity
Mutation
Rare and often linked to diseases
Know how SNPs are used in GWAS studies
GWAS REVERSE GENETIC APPROACH has 3 different qualities
Observe naturally occurring SNPs and mutations
The approach used in humans
Sequence to determine molecular cause
Be able to identify/contrast candidate gene approaches with genome-wide approaches
Candidate/Pathway Gene Association Studies
Penetrance of genetic risk factor
Look for SNPs
GWAS (Genome-wide association studies)
Penetrance of genetic risk factor
Linkage approach and Association approach
No knowledge of what genes may be involved, so they look at the entire genome for genetic variation to identify ASSOCIATIONS with genotype to phenotype
How might one approach be used to inform the other?
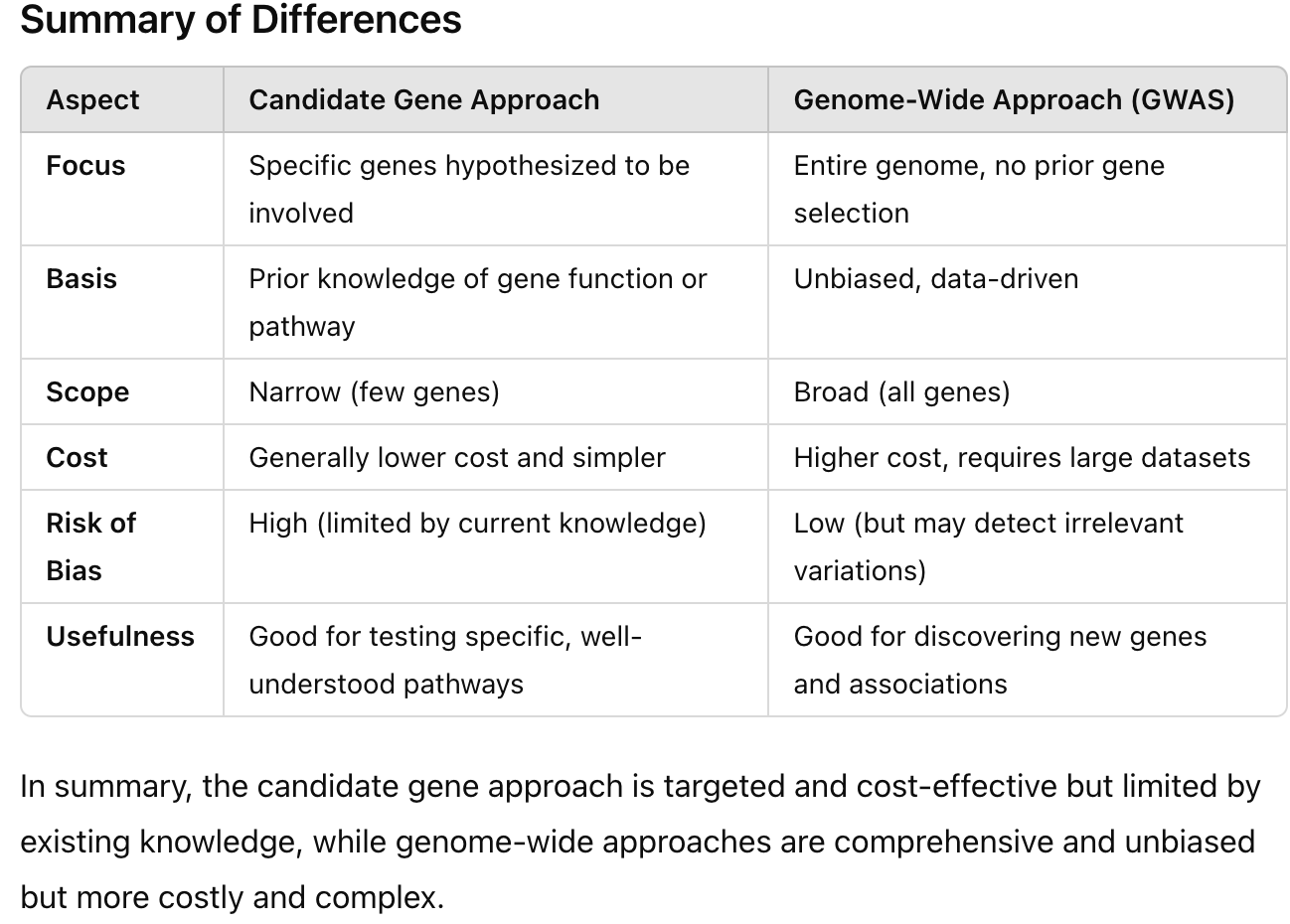
Be able to compare/contrast the two genome association study perspectives.
GWAS (Genome-wide association studies)
Penetrance of genetic risk factor
Using LINKAGE ANALYSIS (1)
Evidence is sought for co-segregation between a LOCUS and putative disease locus using family data
Crossover in chromosomes
Using ASSOCIATION approach (2)
Evidence is sought for an association between a particular ALLELE and disease in a POPULATION
Done in unrelated indivisuals
Powerful for detecting loci with smaller effects
Fine mapping
Doesn’t require family data which means it is less costly and faster
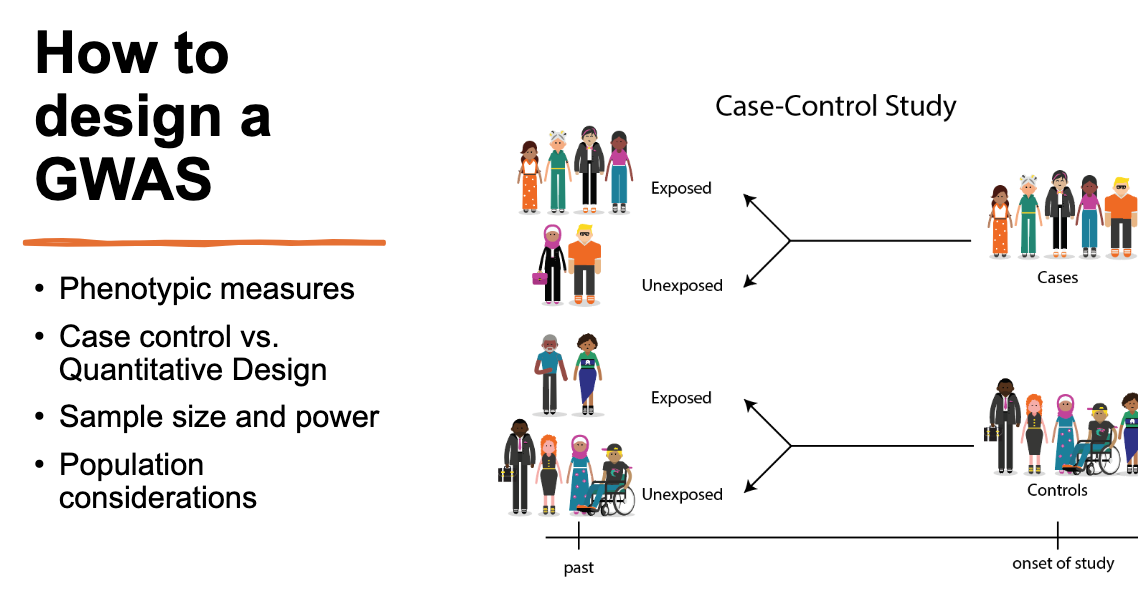
Know the steps of study design, especially population recruitment.
Null hypothesis = There is no association between disease x and marker y
Alternative hypothesis = There is an association between disease x and marker y
Be able to read a Manhattan Plot.
Used in GWAS
Displays genetic variants and a trait (genotype associated with phenotype)
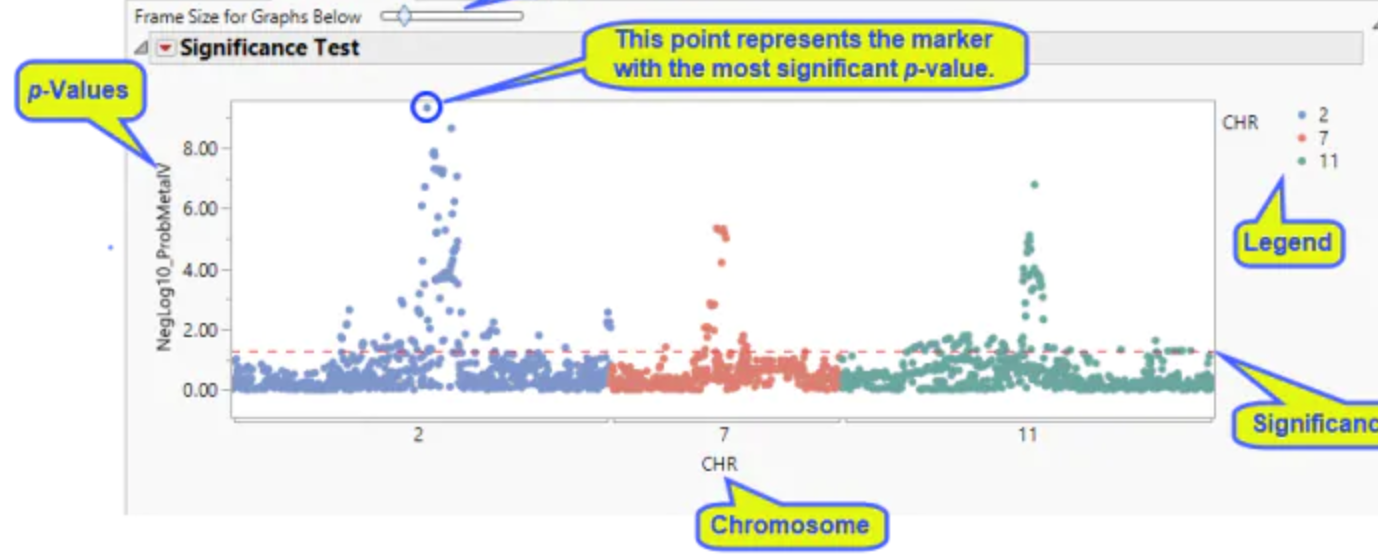
So chromosomes with several points well above the significance threshold line suggest that there may be one of more genes on that chromosome that a strongly associated with the trait being studied.
Type of big question I might ask:
in terms of Sequencing and GWAS: Person with unknown disease, what strategy would you use to determine what gene is mutated? More broadly, you have a population with a disease, how would you design a study?
The reverse genetics approach of GWAS is used in humans to observe the naturally occurring SNP or mutation and then sequence to determine the molecular cause. GWAS also is heavily used in trying to understand SNPs and mutations of genotype and how it is associated with phenotype in populations. In GWAS there are two approaches, linkage and association. Since there is a population with a disease I would use the association approach because the association approach is looking for an association between a particular allele and disease in a population. This tends to be used after linkage, but this is the main approach I would say is most important. By doing this I could compare patient DNA to non-patient DNA of disease-specific SNPs and non-disease SNPs. This would allow me to evaluate the specific person or candidate’s gene(s). In terms of sequencing, I would use next-generation sequencing (NGS) because it allows for larger DNA sequencing to occur. I would use NGS to get larger and more sensitive readings and quantitative results faster. This is all considering the one person is also tied to a population.
Aneuploidy
What makes a chromosome a chromosome? (in DNA repair lecture)
What is a Karyotype? What does it measure?
A person’s complete set of chromosomes
Know some examples of Aneuploidy. Cancer, Down’s syndrome
Aneuploidy = change in the number of chromosomes in the genome
Not all aneuploidy = death
Ex. sex chromosomes, down syndrome
Cancer – 26 chromosomes in healthy human cell, 59 chromosomes in colorectal cancer cell
Why is Aneuploidy bad?
Death, health issues
How does Aneuploidy happen?
Nondisjunction
What are the major chromosomal rearrangements.
Use the examples to help your understanding but you won’t be directly tested on them.
Single Chromosome Structural Changes
Deletion
loss
Duplication
Gain
Inversion
Two Chromosome Structural Changes
Insertion
Translocation
In terms of viability, a balanced translocation could result in how many live births? Why?
2 live births out of 4 outcomes
Normal
Balanced translocation
Unbalanced translocation
Severely unbalanced translocation
two outcomes (normal karyotype and balanced translocation) are typically viable and can result in live births. Therefore, in terms of viability, a balanced translocation in a parent generally results in about 50% of gametes that can potentially lead to live births.
How can you experimentally find chromosomal abnormalities?
Fluorescence In Situ Hybridization
Chromosome microarray
Next Generation Sequencing
Linkage analysis and genome sequencing
How do chromosomal rearrangements happen?
Chromosome breakage
Problems with DNA repair
Exogenous (outside DNA that is introduced into its cells) DNA-damaging agents
What is a common fragile site?
CENTROMERE
METAPHASE
NON-DIVIDING
Type of big question I might ask:
in terms of Aneuploidy: I might give you a karyotype and ask you to determine the rearrangement and predict some of the consequences in terms of structure or function.
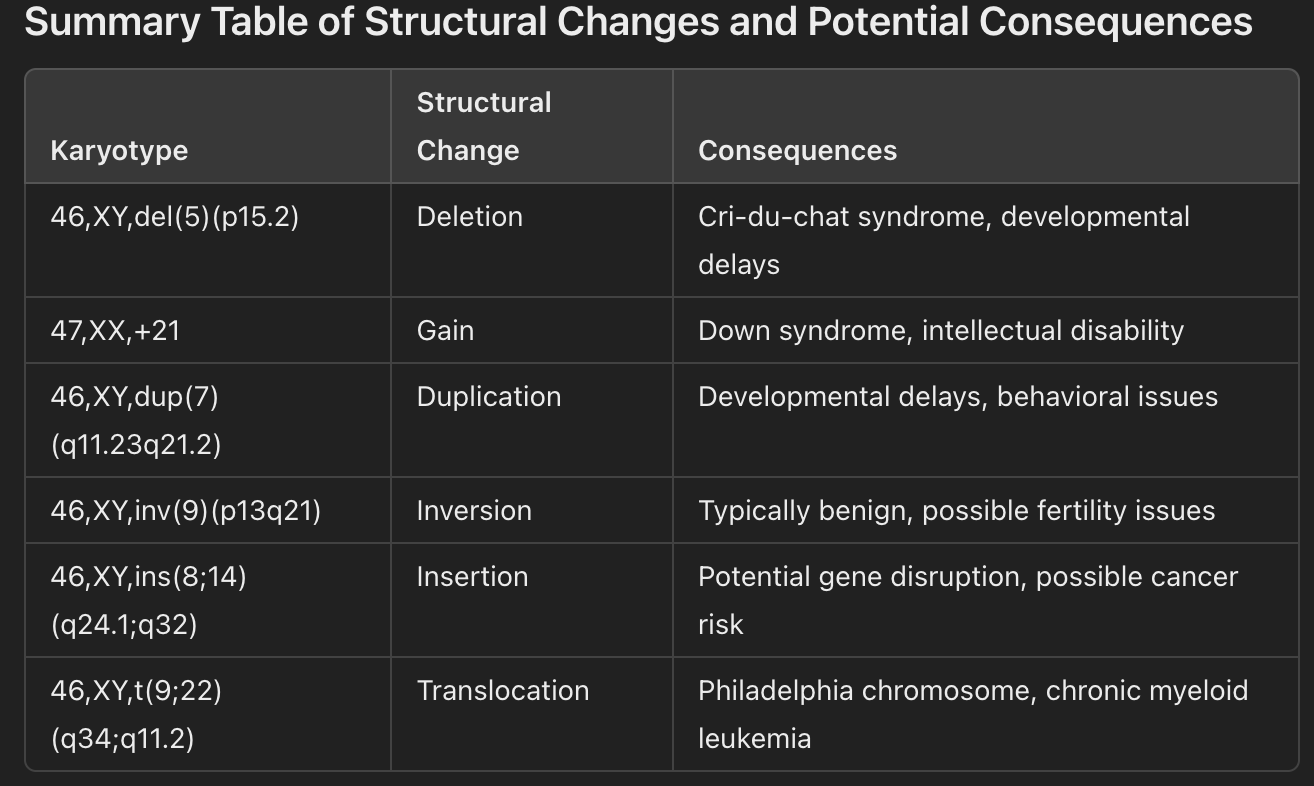
1. Deletion (Single Chromosome Structural Change)
Karyotype Notation: 46,XY,del(5)(p15.2)
Description: This indicates a deletion on chromosome 5 at the short arm (p) in the region p15.2.
Consequences:
The loss of genetic material in this region can disrupt normal development and lead to Cri-du-chat syndrome, which is associated with intellectual disability and developmental delays.
Loss of function in genes located in the deleted segment could result in a loss of protein functions important for neurological development and other bodily functions.
2. Gain (Aneuploidy)
Karyotype Notation: 47,XX,+21
Description: This represents a gain of an entire chromosome (trisomy 21), meaning there are three copies of chromosome 21.
Consequences:
Trisomy 21 causes Down syndrome, characterized by intellectual disability, distinct facial features, and increased risk of congenital heart defects.
Having an extra copy of chromosome 21 means an overexpression of the genes on this chromosome, leading to structural and functional abnormalities across various systems.
3. Duplication (Single Chromosome Structural Change)
Karyotype Notation: 46,XY,dup(7)(q11.23q21.2)
Description: There is a duplication on chromosome 7, specifically on the long arm (q) between bands q11.23 and q21.2.
Consequences:
A duplication can result in overexpression of genes within this region, potentially affecting cognitive function, growth, or physical development.
This specific duplication is linked to developmental and speech delays and can sometimes result in behavioral issues.
4. Inversion (Single Chromosome Structural Change)
Karyotype Notation: 46,XY,inv(9)(p13q21)
Description: This indicates an inversion on chromosome 9, where the segment between bands p13 and q21 is flipped.
Consequences:
Inversions generally don’t cause issues unless they disrupt specific genes, though they may lead to fertility issues due to complications in chromosome pairing during meiosis.
If genes within the inversion are disrupted, it may lead to abnormal gene function, potentially causing developmental or health issues.
5. Insertion (Two Chromosome Structural Change)
Karyotype Notation: 46,XY,ins(8;14)(q24.1;q32)
Description: This indicates an insertion of genetic material from chromosome 14 (band q32) into chromosome 8 at q24.1.
Consequences:
This insertion could disrupt the function of genes at both insertion sites, potentially leading to structural or functional abnormalities depending on the genes involved.
Insertions in this region are sometimes associated with certain types of cancer due to disruption of oncogenes or tumor suppressor genes.
6. Translocation (Two Chromosome Structural Change)
Karyotype Notation: 46,XY,t(9;22)(q34;q11.2)
Description: This indicates a translocation between chromosomes 9 and 22, with segments from q34 on chromosome 9 and q11.2 on chromosome 22 swapping places.
Consequences:
This specific translocation forms the Philadelphia chromosome, commonly associated with chronic myeloid leukemia (CML).
The fusion of the BCR gene on chromosome 22 and the ABL gene on chromosome 9 creates the BCR-ABL fusion gene, leading to unregulated cell division and cancer.
DNA repair
Know the kinds of mutations.
Repeat expansions
Point mutations
A single nucleotide base in the DNA sequence is altered
Missense
A single nucleotide base change leads to a different amino acid in the protein
May alter protein function
Nonsense
A single nucleotide base change creates a premature stop codon
Nonfunctional protein is the result
Silent
Change in a nucleotide base that does not alter the amino acid
Frameshift mutations
Insertion or deletion of nucleotides that change the reading frame for the gene
Since reading nucleotides occurs in triplets (the codon) adding or removing not in multiples of 3 alters every other codon
Usually nonfunctional protein
Insertion mutations
One or more nucleotides are added to the DNA sequence
Can disrupt gene function if it causes a frameshift or adds extra amino acids to the protein
Deletion mutations
One or more nucleotides are removed from the DNA sequence
Can disrupt gene function if it causes a frameshift or results in loss of essential parts of the protein which can impair its function
Copy Number Variations (CNVs)
Large segments of DNA are duplicated or deleted
Repeat expansion mutations
A sequence of DNA bases that normally repeats a few times is expanded to repeat many more times
Often trinucleotide (3 bases) repeats
CAG, CGG, or GAA
Inversion mutations
A segment of DNA is reversed within the chromosome
Translocation
A segment of DNA is moved from one chromosome to another
Know the common causes of DNA damage, types of DNA lesions
Base mismatch
Single-strand break
Double-strand break
Interstrand corsslinks
Bulky adducts/ intrastrand crosslinks
Events that can lead to DNA breakage
Exogenous (outside DNA being introduced to cells) DNA-damaging events
Problems with replication or DNA repair
Barriers to replication
Replication fork barriers
DNA damage
protein/DNA interactions
DNA secondary structures
RNA: DNA hybrids
DNA repair related to DNA replication/DNA base damage
Mismatch repair
Mis-incorporated bases
Base excision repair
Damaged bases or uracil incorporation
Nucleotide excision repair
Removal and repair of bulky damage
Know the two kinds of double-strand break repair
Know the major steps of NHEJ and HR
Non-Homologous DNA
DNA sequences that lack a significant similarity and do not share a recent common ancestry
No sequence similarity
Unrelated genes or chromosomal regions
Two different chromosomes
Genes with distance functions
Homologous DNA
DNA sequences that have similar or identical sequences due to shared ancestry
Same gene or chromosome
Genes in related species
Sister chromatids (created after DNA replication in the S phase)
Non-homologous end-joining (NHEJ)
Does not need a template – directly joins the broken DNA ends
Aligning broken ends
Proteins bind to the broken ends
End processing
Trimming or filling in the nucleotides
Ligation
Ends are joined together
Homologous Recombination
Uses homologous DNA (like sister chromatids) as a templete
Homologous template
Resection (generation of ssDNA tails)
Homology search and invasion
ssDNA gaps are filled in
What is a Holliday Junction and a cross-over?
DSBR (Double-Strand Break Repair)
Single strand invasion
Second end capture
HJ
Resolution
Crossover
Either non-crossover or crossover
Type of big question I might ask in terms of DNA repair
I might give you a scenario where there is a mutation in a DNA repair protein, and you’ll have to think about some of the consequences of that. Alternatively, I might create a type of DNA lesion and ask you to determine what pathway is likely involved in repairing it.
Scenario 1: Mutation in a DNA Repair Protein
Imagine there’s a mutation in the BRCA1 gene, which encodes a protein involved in the homologous recombination (HR) repair pathway. The mutation disrupts the function of the BRCA1 protein, making it unable to effectively bind to other proteins involved in HR.
Question:
What might be the consequences of this mutation for the cell?
BRCA1 mutations would disrupt the protein's ability in HR which is crucial for repairing DSBs (doub-stranded breaks). This could cause the cell to resort to NHEJ (non-homologous end-joining) which is quicker but is more error-prone.
Errors can mean mutations and chromosomal reaaragments
Mutations:
Point
Frameshift
Insertion
Deletion
CNV
Repeat
Inversion
Translocation
Chromosomal rearrangements:
Single Chromosome Structural Changes
Deletion
Duplication
Inversion
Two Chromosome Structural Changes
Insertion
Translocation
How could this impact the cell’s ability to repair DNA double-strand breaks, particularly during the S and G2 phases of the cell cycle?
This forces the cell to rely more on NHEJ, which could lead to mutations during DNA replication in the S phase, potentially leading to disrupted cell function or cell death.
What are some potential long-term effects of impaired BRCA1 function on cell health or cancer risk?
Increased Cancer Risk: Cells with BRCA1 mutations accumulate DNA damage over time, which can activate oncogenes or inactivate tumor suppressor genes, leading to cancer. This is why BRCA1 mutations are closely linked with breast and ovarian cancers.
Early Cell Death or Senescence: Accumulated DNA damage can make cells more likely to enter senescence (a non-dividing, “aged” state) or undergo apoptosis (programmed cell death), impacting tissue health and regeneration.
Scenario 2: DNA Lesion and Repair Pathway
Imagine a cell has been exposed to UV light, causing pyrimidine dimers to form in its DNA. These dimers create covalent bonds between adjacent thymine or cytosine bases, distorting the DNA helix.
Question:
Which repair pathway is most likely to be involved in fixing this type of damage?
What are the basic steps of this repair pathway, and what are some key proteins involved?
What might happen if this repair pathway is defective or overwhelmed?
Organellar inheritance
Know that chloroplasts and mitochondria are the two organelles that maintain their own genome.
Know the basics of the mitochondrial genome—what kinds of genes are in the genome.
Be cognizant of the fact that the electron transport chain uses proteins encoded by both the nuclear and mitochondrial genomes
Circular chromosome
Heavy (Guanine rich)
Light (Citpcine rich)
Many copies in each cell
Contains 37 genes
Lack of histones
Maternal inheritance
Multiple copies per cell
Be able to identify a pedigree that is indicative of organellar inheritance(especially compared to a Mendelian inheritance pedigree)
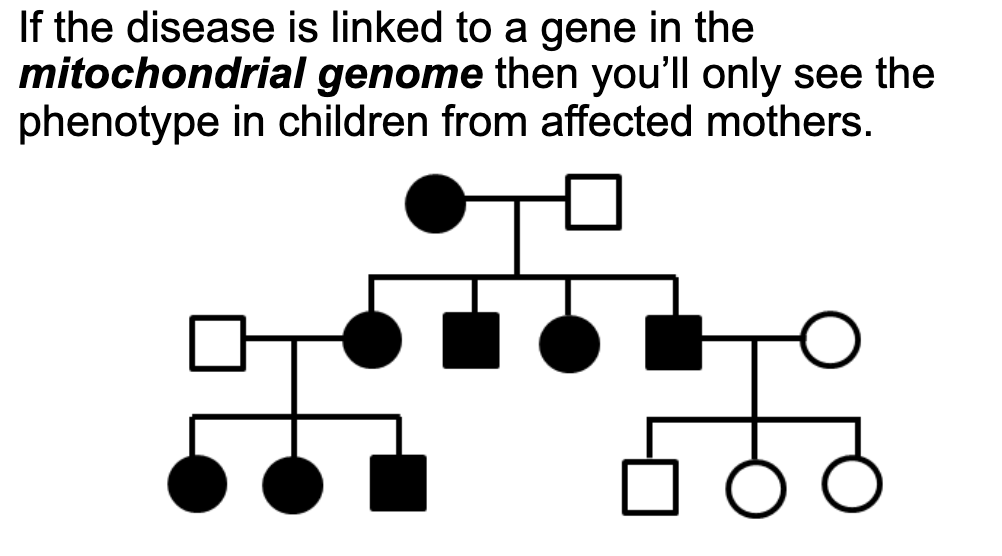
Understand that the mitochondrial genome is more likely to accumulate mutations
Further, due to random mitochondrial selection in oogenesis, there may be variable penetrance in genetic disorders.
Extra nuclear inhertienace = non mendelian inheritance
DNA in plicating mitochondria and some other organelles of the cell
Generally only one parent ccontributes
Type of big question I might ask:
Compare and contrast a pedigree, what is the inheritance pattern? The children all have variable phenotypes, why?
Journal Club: Know what a genetic assay is and how the YAC assay works generally.
Yeast Artificial Chromosome
Genetically engineered vector made from yeast DNA
Allows researchers to clone and analayze large DNA sequences
Used for
Genome mapping
Study gene regulation
General Steps in How YACs Work:
Preparation of Vector Components: The YAC vector contains essential yeast DNA elements like centromeres (CEN), telomeres (TEL), and autonomously replicating sequences (ARS) to maintain stability and replicate like yeast chromosomes.
Inserting the DNA Fragment: A large DNA fragment from the organism being studied is inserted into the YAC. This DNA fragment contains the target gene or region researchers want to study.
Transformation into Yeast: The YAC with the inserted DNA fragment is then introduced into yeast cells. Since yeast cells recognize the YAC as a regular chromosome, they replicate and segregate it during cell division.
Studying and Analyzing the DNA: Once in the yeast cells, the inserted DNA can be studied as though it’s in a chromosome. Researchers can perform genetic assays to observe the behavior, expression, or structure of the inserted gene.
Genetic Assay
Lab technique used to study and measure specific genetic sequences, gene expression, or genetic changes
Identify mutations
PCR
qPCR
Gel electrophoresis
Genes and Genomes
Unit 2- Exam
Past questions from old exams
Note, these questions are across an upper - level molecular biology online course (think
Molecular and Cellular Bio with Dr. Kwan) and a molecular genetics lab that was specialized on
chromosome fragility (Fra16D). Not all situations have been replicated in this course, but this might give you some sense of some basic questions I might ask that is related to content in this unit.
True/False:
A chromosome is defined by having one telomere, a centromere, and an origin of replication
FALSE
Fragility is a measure of DNA repair.
FALSE
Hemizygous and homozygous deletions at common fragile sites are more commonly found in cancers.
TRUE
End-point PCR is quantitative
FALSE
A transition mutations is replacement of a Purine:Pyrimidine base pair with a Pyrimidine: Purine base pair, or vice versa.
FALSE
Transition mutations are purpine substituted with purine ( A - G ) or pyrimidine substituted with pyrimadine ( C - T )
A transversion mutation is replacement of a Purine:Pyrimidine base pair with a Purine:Pyrimidine base pair, or vice versa.
TRUE
Fill in the blank
Recombination/Replication is the process by which a genome is copied in cells.
Base excision repair recognizes small (type of lesion).
Nucleotide excision repair recognizes bulky (Pyrimadine dimers) (types of lesion).
Multiple Choice
What DNA structure is predicted to form at long tracts of AT repeats:
G quadruplex
Hairpins
Cruiciforms
Triplex
No structure
11) Where is FRA16D normally located?
In an oncogene
In a tumor suppressor gene
Near the telomere
On a human artificial chromosome
In the mitochondrial genome
12) What is the molecule that is used as a chain terminator in Sanger sequencing and base pairs with guanine?
dideoxycytosine triphosphate
cytosine nucleotriphosphate
cytosine triphosphate
cytosine triphosphate
deoxycytosine triphosphate
13) Indels:
Are always silent mutations
Can result in a frameshift
Are the insertion of a premature stop codon
Are due to transition mutations
14) Synthesis dependent strand annealing (SDSA) will result in:
One Holliday junction, no crossing over
One Holliday junction, non-reciprocal crossing over
Two Holliday junctions, crossing over
Two Holliday junctions, no crossing over
14) Which of the following substrates is likely to be repaired by HR?
A broken replication fork
A bulky DNA adduct
Single strand breaks
A mismatched DNA base
15) I am a Physician Scientist and have a patient with a family history of melanoma (skin
cancer). I decide to enroll them in your study and do whole genome sequencing on their skin cells. Excitingly you find a novel SNP that is predicted to end in a premature stop codon of a known member of a DNA repair pathway. For this patient, what DNA repair pathway are they likely deficient in?
Base - excision repair
Nucleotide - excision repair
the DNA repair pathway primarily responsible for repairing bulky, helix-distorting lesions, such as those caused by UV radiation (e.g., thymine dimers).
Mismatch Repair
Non - homologous end - joining
Homologous recombination
*My intention was for this to be NER (melanoma being the key term to signal
NER deficiency) but a lot students chose MMR, that I allowed that to also be
an acceptable answer.
16) Extended treatment with the drug hydroxyurea results in replication fork stalling and
eventually replication fork collapse. What DNA repair pathway is important in fork restart at a collapsed fork?
Base-excision repair
Nucleotide-excision repair
Mismatch Repair
Non -homologous end-joining
Homologous recombination
HR is involved in repairing double-strand breaks (DSBs) that occur at collapsed replication forks by using an undamaged homologous template (such as the sister chromatid) to accurately restore the DNA sequence.
Experimental questions:
17) Consider the Homologous recombination. Put the pathway in order.
Homologous template
Resection (generation of ssDNA tails)
Homology search and invasion
ssDNA gaps are filled in
18) 5’ATGGCGACCCTGGAAAAGCTGATGAAGGCCTTCGAGTCCCTCAAGTCCTC
CAGCAGTTCCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAGCAACAGCCGCCACCGCCGCCGCCGCCGCCGCCGCCTCCTCAGCTTCCTCAGCCGCCGCCG 3’
The above is a region of DNA that contains a CAG repeat tract. Why would it be bad
idea to design your primers within the CAG repeat tract itself?
Unstable primer binding
19) This above DNA strand is GC rich. DNA must be denatured before synthesis can
occur. What might you need to take into consideration regarding denaturation when
designing your PCR program for the CAG repeat?
Gc rich regions have stronger base pairing since G-C is bonded by 3 hydrogen bonds.
Heat must be higher to denature or break the bonds
20) What would happen if you forgot to add dideoxycytosine triphosphate (ddCTP) and
only added a limiting amount of Deoxycytidine triphosphate (dCTP) to your Sanger
sequencing reaction (4 pts)?
The sequencing process would fail to properly terminate resulting in non-terminated fragments
21) Name one potential barrier to replication (2 pts)
DNA secondary structures can form due to sequences rich in specific bases like the one from above. They can create G-quadruplexes which can physically block the progression of the replication fork preventing DNA sysnthesis
Other secondary strcutures like the ahirpin and cruciform can and cause the replication fork to break which would call for homolgous recombination to repair it.
Word Problem:
22)
You’re a student researcher studying a novel Fragile sequence, Fra17A. You have a set of genomic DNA samples from various individuals and find that Fra17A has a polymorphic CGG repeat like that seen in Fragile X syndrome, a rare fragile site. As part of your project, you clone varying lengths of the CGG rep eat into our YAC assay. You successfully make a CGG0, CGG10, CGG40 and CGG 80 YAC strains
(Wow! That was hard work!). You do a fragility assay and measure FOA resistance
and find the following relationship:
23) As part of your project, you decide to delete Rad52, a protein essential for repair of double strand breaks. Based on your knowledge of DNA repair and fragility, draw a figure showing your expectations of what a RAD52 mutant will do to fragility compared to wildtype.
24) Next, to wildtype cells with the CGG repeat YAC, you add a drug that impairs DNA replication (hydroxyurea). Now draw a figure showing your expectations of what a stalling replication will do to fragility compared to wildtype cells treated with no drug.
25) What experimental method might you use to measure whether the replication fork stalls at the CGG repeat?
I would use qPCR since it can be used to measure the amount of DNA at different time point. You are getting real time readings so I would be able to observes any delayed or reduced replication at the CGG repeat and it can be quantified as well.