Individual genotype: Individual Identity and Capture Mark Recapture by Genotype
Marker: a gene or DNA sequence that allows us to detect variation among individuals or between alleles
Brief overview of Mendelian inheritance
all cells in mammals and birds, except sperm, eggs, and RBC in mammals, are diploid (they have two copies of each type of chromosome)
Chromosome: a distinct piece of DNA (and associated proteins) containing many genes (23 dif chromosomes in humans, 2n=46)
you inherit one copy of each chromosome from your mother and one from your father, therefore you have two alleles at every locus
Allele: genetic variant or alternative form of a gene (think yellow or green in peas)
Genotype: the specific alleles present in an individual at a particular locus or many loci (or the entire genetic makeup of an individual)
Markers: need to be variable
common types of variable DNA markers
SNPs
Microsatellites
DNA sequences and AFLPs (will cover later on)
RFLPs and many others that we won’t discuss
SNPs
Single nucleotide polymorphism: single base pair positions that vary among individuals (PCR-based)
Genetically characterize individuals and populations
SNPs plentiful: every 200-500 bp for non-coding DNA and every 500-1000 bp for coding DNA
SNPs are becoming more common bc of new methodological approaches, can obtain 1000s of SNP markers - very powerful
Microsatellites (msats)
diploid marker
Short Repeats (ex. ATATATATATAT)
present in most organisms and highly variable: can detect differences among pops or closely related individuals
Fast mutation rates: can look at short-term changes (not good for changes in the distant past)
Neutral “junk” DNA
Advantages: PCR-based so almost any tissue can be used. Tiny amounts of tissue required
Disadvantages: Primer of known sequence required. Can be expensive and time-consuming to develop
probably most common approach for conservation genetics
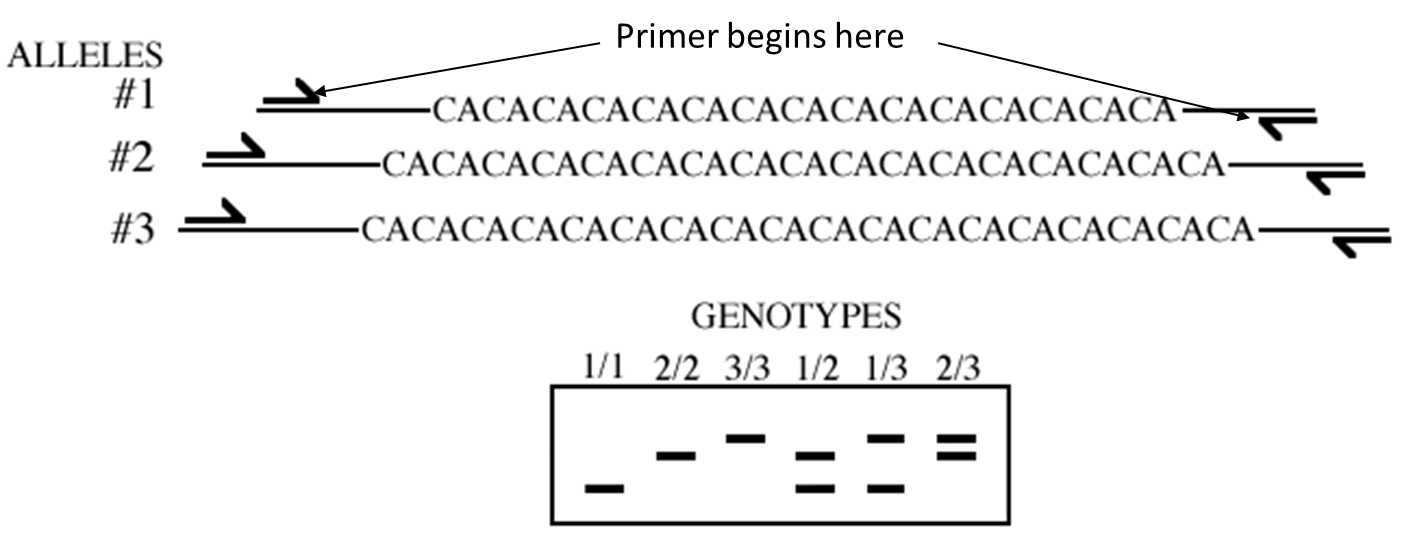
One Locus
Homozygous vs heteroxygous
Examples of microsatellite data
to estimate Nc (census size)
to assess parentage/familial relationships
Using genetics to estimate Nc
number of individuals in a pop is a fundamental characteristic
why not use traditional methods?
can use genetics to estimate Nc
rarefaction
capture-mark-recapture
Rarefaction
estimate minimum pop size by counting the number of unique genotypes
ex. 30 unique genotypes out of 115 coyote feces samples
estimate can be corrected to take into account the probability of not sampling individuals
Capture-mark-recapture - Individual ID: Match probability
match the genotype of a sample to an individual
ex. match a hair to an individual bear to recognize a recapture or a new individual
use highly polymorphic markers and then compute a match probability based on allele frequencies from the reference pop
Capture-mark-recapture
can be used with genetic data
Multilocus genotypes are the permanent marks
Nc = (N1 x N2)/R
N1 = number of individuals int he first sample
N2 = the number of individuals in the second sample
R = number of animals recaptured
Ex: 10 animals caught in first, 10 caught in second, 5 animals capture in the second sample were marked
Nc = (10×10)/5 = 20
Problems with the method
failure to distinguish individuals (not enough loci, loci not variable enough) = underestimate pop size
Genotyping errors produce false unique genotypes = overestimate pop size
Bears - Northern Continental Divide Ecosystem
more sophisticated: not just estimating abundance but long-term monitoring of pop growth rates (lambda) and distribution
Grizzly Bears
listed under the ESA since 1975
Monitoring consisted of opportunistic sightings of females with cubs, distribution of females with young, and human-caused mortalities
Inadequate: no reliable estimates of pop size, trend, or distribution
2006: Montana draft grizzly bear management plan: pop trend info will guide management decisions
Radiotrack 25 females in perpetuity
Live capture of bears is:
expensive
logistically difficult in remote areas
requires specialized training
has inherent risk to bears and trappers
may be subject to intense scrutiny and potential moratoria on public lands
Non-invasive genetic sampling
barbed wire on bear rubs
Species ID, individual identity, and sex via nuclear DNA extracted from hair follicles
7 msats
Objectives
power to detect gender-specific and pop-wide declines in pop abundance
precision and relative bias of growth rate estimates
Sampling effort required to achieve 80% power to detect a decline within 10 years
Results
model simulations showed annual bear rub surveys would exceed 80% power to detect 3% annual decline within 6 years
rate of decline rapid enough to require management intervention, but slight enough to require a powerful monitoring method
Parentage Analysis
can use individual genotypes to identify mothers and fathers bc we know that one allele at each locus comes from the mother with the other allele comes from the father
can do with a specific group of individuals
ex. nestlings where the mother is known
or for a pop without a pedigree
here you need many and highly variable loci
can assign probable parents for each individual and build a pedigree