Microbiology week 3 LSINT
Unicellular Eukaryotes and Protists
Unicellular eukaryotes (excluding yeast and algae) are very diverse.
Many are parasites.
Examples mentioned: Diatoms, Oomycetes (water molds), Amoebozoa, Slime molds.
Concept: Protists as a broad, diverse group within eukaryotes.
Visual aids referenced (images of protozoan-like organisms).
Fungi: Yeasts vs. Molds
Yeasts: unicellular; grow by cell division.
Molds/filamentous fungi: multicellular; grow as filaments.
Mycelium: network of hyphae.
Aerial hyphae: hyphae that extend into the air.
Production of spores (asexual) as part of fungal life cycles.
Overall note: Fungi include yeasts and molds with distinct growth forms.
Fungal Cell Wall and Immunology
Fungal cell walls are recognized by the host immune system.
Reference: Hardison, S.E. & Brown, G.D. (2012) C-type lectin receptors orchestrate antifungal immunity. Nature Immunology 13, 817–822. DOI: 10.1038/ni.2369
Implication: Fungal pathogens interact with innate immune receptors (e.g., C-type lectin receptors) that shape antifungal responses.
Bacterial Cell Walls: Gram-Positive vs Gram-Negative
Gram-negative bacteria:
Outer membrane present.
Periplasmic space.
Peptidoglycan layer.
Porins and lipopolysaccharide (LPS) in the outer membrane.
Gram-positive bacteria:
Thick peptidoglycan layer.
Teichoic acids and lipoteichoic acids.
Lipoproteins associated with the cell membrane.
Key contrast: Outer membrane and periplasm vs thick peptidoglycan layer without outer membrane.
Peptidoglycan Synthesis in Bacteria
Peptidoglycan synthesis involves:
Transglycosylase activity at growing point.
Cytoplasmic membrane involvement; translocation of precursors (outward/inward movement).
Pentapeptide subunits: D-Ala–D-Ala, L-Ays, DAP (diaminopimetic acid) variants; specific cross-links via transpeptidation.
Bactoprenol serves as the lipid carrier for peptidoglycan precursors.
Autolysins help remodel the cell wall during growth.
Process outline (as depicted in Brock Fig. 5.8):
Growing point at the cytoplasmic membrane.
Transglycosylation inserts peptidoglycan subunits into the cell wall.
Transpeptidation cross-links occur to form mature wall.
D-Ala and related residues cycle as part of the cross-linking process.
Significance: Peptidoglycan architecture is a critical target for antibiotics (e.g., β-lactams) and differentiates Gram categories.
Key Questions for Viruses (Overview from slides)
1) What is a virus / virion?
2) Size of viruses.
3) Baltimore classification (genome and replication strategy).
4) Virus replication cycle in bacteria and animal cells.
5) Lytic vs lysogenic pathways in bacteria.
6) Outcomes of viral infection in animal cells.
7) Bacterial defenses against viral infections (e.g., CRISPR).
Note: Viral topics of current interest include comparisons between viruses infecting bacteria vs. animal cells.
Importance and Roles of Viruses
Viruses cause disease in humans, animals, and plants (in the context of medical microbiology).
Viruses as tools in molecular biology/biotechnology: gene insertion/expression systems (proteins)
Use as therapy: bacteriophages used to prevent or treat infections (phage therapy).
Obligate Intracellular Parasites
Viruses require cellular hosts for energy and protein synthesis.
They infect plant cells, animal cells, and even bacterial cells.
Virion = infectious particle.
Example: Tobacco Mosaic Virus (TMV) discovered by Martinus Beijerinck in 1898.
Concept: Virus–host interactions depend on the host cell machinery; viruses are inert outside a host.
Tobacco Mosaic Virus (TMV)
TMV is a classic example of an RNA virus with a rod-shaped virion; historically important for early virology discoveries (Beijerinck, 1898).
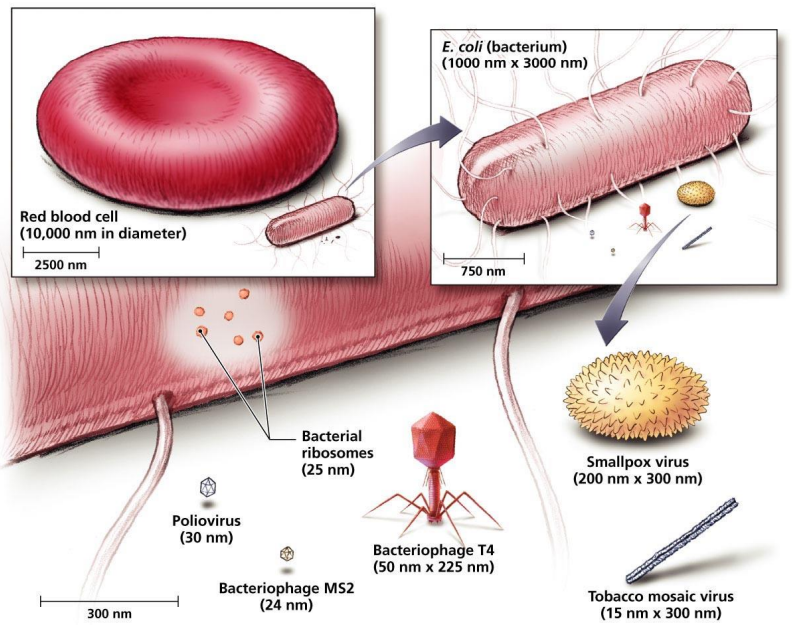
Size Scales: Cells, Bacteria, and Viruses
Red blood cell diameter: approximately 10{,}000\mu{m} (i.e., 10 μm).
E. coli bacterium: approximately 1{,}000 ext{ nm} imes 3{,}000 ext{ nm} .
Bacterial ribosomes: approximately 25 ext{ nm} in diameter.
Smallpox virus: approximately 200 ext{ nm} imes 300 ext{ nm}.
Poliovirus: approximately 30 ext{ nm}.
Bacteriophage MS2: approximately 24 ext{ nm}.
Bacteriophage T4: approximately 50 ext{ nm} imes 225 ext{ nm}.
Tobacco mosaic virus (TMV): approximately 15 ext{ nm} imes 300 ext{ nm}.
General note: viruses span from tens to a couple hundred nanometers; some giant viruses exceed 300 nm and blur traditional size boundaries.
General size range: approximately 0.02 ext{ μm} ext{ to } 0.3 ext{ μm} for many viruses (with larger exceptions like giant viruses).
Viral Genome and Virion Structure
Viral genome characteristics:
DNA or RNA genome.
Single-stranded (ss) or double-stranded (ds).
Linear or circular forms.
Genome packaging:
Genome + capsid = nucleocapsid.
Capsid built from capsomeres (protein subunits).
Viral envelope:
Some virions are enveloped, with a membranous envelope surrounding the capsid.
Virion components:
Capsid: helical or icosahedral symmetry; composed of capsomeres.
Genome inside capsid; sometimes with envelope and, in some cases, tail structures (e.g., bacteriophages).
Representative virion families (examples shown in figures):
Tobacco mosaic virus (TMV)
Adenoviruses
Influenza viruses
Bacteriophage T4
Overall takeaway: Virions protect the genome and aid in host cell entry; the genome can be DNA or RNA, with various configurations.
Baltimore Classification: Seven Classes
Class I: dsDNA (+/-) viruses; examples include Herpesviruses.
Class II: ssDNA (+) viruses; example Parvovirus.
Class III: dsRNA (+/-) viruses; example Reovirus (Rotavirus).
Class IV: ssRNA (+) viruses; example Coronaviruses, Poliovirus (replication to produce genome RNA and mRNA).
Class V: ssRNA (-) viruses; example Influenza virus, Rabies virus.
Class VI: ssRNA (+) retroviruses; example HIV; replication involves reverse transcription.
Class VII: dsDNA viruses that replicate through a reverse-transcribed RNA intermediate; example Hepatitis B virus.
Concept: Baltimore classification groups viruses by genome type and replication strategy, not by host range or morphology.
DNA and RNA Virus Replication: Synthesis of Viral mRNA
DNA viruses(host RNA polymerase–driven transcription into mRNA):
Class I & VII use host machinery for mRNA synthesis from DNA genomes.
dsDNA (+/-) viruses (e.g., Herpesviruses) and dsDNA viruses that involve reverse transcription (Class VII).
Transcription produces mRNA that is translated by host ribosomes; DNA replication may involve a dsDNA intermediate or other forms.
RNA viruses rely on viral RNA-dependent RNA polymerase to generate mRNA from genomes:
Class III dsRNA, Class IV (+) RNA, Class V (-) RNA, Class VI (+) RNA retroviruses, Class VII (dsDNA with reverse transcription) uses reverse transcription during replication (but the mRNA synthesis step is described under DNA virus discussion).
Illustrative mapping (from Figure 11.2 style content):
DNA viruses: Host RNA polymerase converts viral DNA to mRNA; replication involves dsDNA and possible replicative forms.
RNA viruses: Viral RNA replicase produces mRNA and negative/positive strand intermediates as required by class.
Significance: Different replication strategies impact antiviral targeting and gene expression strategies.
DNA vs RNA Viruses: Synthesis of Viral mRNA (Detailed View)
DNA viruses (host RNA polymerase –> mRNA):
dsDNA (+/-) viruses use host RNA polymerase for mRNA synthesis.
Examples: Phage T4 (Class I) and Hepatitis B (Class VII – reverse transcription stage).
Transcription produces mRNA from the viral genome; a dsDNA intermediate (replicative form) may be involved for some classes.
RNA viruses (viral RNA replicase–mediated):
Class III: dsRNA, Class IV: (+) RNA, Class V: (-) RNA, Class VI: (+) RNA retroviruses, Class VII: dsDNA with reverse transcription (as noted).
Viral RNA replicase synthesizes mRNA from the viral RNA genome; translation by host ribosomes then occurs.
Baltimore classification emphasizes how the genome type dictates replication and transcription strategy.
Viral Replication Cycle: Steps and Phases (General)
Core steps of the replication cycle (bacterial virus example):
Attachment/adsorption to host cell.
Penetration/entry (injection of genome in the bacterial cell).
DNA replication and transcription/translation in the host.
Synthesis of viral components.
Assembly and packaging of new virions/viro particles.
Release from the host cell (often via lysis in bacteriophages).
Phases:
Eclipse phase: period during which no infectious virions are detected outside the cell; viral components are being produced inside.
Maturation phase: newly assembled virions become capable of infection within the host cell.
Release phase: the final stage where mature virions are released from the host cell, which can occur through lysis or budding, allowing them to be seen or infect new host cells.
Culturing and Growth of Viruses
Host cells required to culture viruses:
Bacteria (bacteriophages).
Animal cells.
Plant cells.
Practical note: Viral culture depends on a permissive host system that supports replication and production of infectious virions.
Temperate Bacteriophages: Lytic vs Lysogenic Pathways
Lytic pathway (virulent infection):
Phage components are synthesized; host cell undergoes lysis; new virions are released.
Lysogenic pathway (temperate phage):
Viral genes are not transcribed initially; genome integrates or persists in host DNA; replication of genome occurs with the host.
The lytic pathway can be reactivated under certain conditions, leading to phage production and cell lysis.
Diagrammatic reference: Brock Fig. 8.16 illustrating attachment, injection of viral DNA, and the two pathways.
Outcomes of Virus Infections in Animal Cells
Classifications of infection outcomes include:
Virulent infection (productive lytic infection in animal cells).
Latent infection (virus genomes persist without immediate production of virions).
Persistent infection (ongoing, low-level production or maintenance).
Transformation (virus-associated cellular transformation and potential oncogenesis).
Reference: Brock Fig. 8.20 describes these outcomes in animal cells.
Viral Infection Lifecycle: A Snapshot of Stages
General lifecycle stages depicted in virus replication figures:
Attachment/Endocytosis/Entry to host cell.
Uncoating to release genome into the cell.
Genome replication and transcription to produce viral RNAs and mRNAs.
Translation of viral proteins by host ribosomes.
Genome replication, assembly of virions in the cytoplasm or nucleus.
Trafficking through cellular organelles (e.g., Golgi apparatus) and exit from the cell.
Visual aids show different organelle involvements (nucleus, ribosomes, Golgi, etc.).
Bacteriophage Infection and CRISPR-Cas Defence
Bacteria possess adaptive immune systems to defend against phage infections:
CRISPR-Cas systems provide sequence-specific defense against invading phages.
Concept: CRISPR-Cas systems capture DNA snippets from phages and use them to recognize future infections, guiding targeted destruction of invader DNA.
Visual: CRISPR-Cas figure illustrating bacterial defense against viruses.
CRISPR-Cas in Biotechnology
Beyond natural defense, CRISPR-Cas systems have been repurposed as powerful biotechnological tools for genome editing and other applications.
Slides reference a dedicated section on CRISPR-Cas in biotechnology, highlighting its broad impact in molecular biology and genetics.
Next Topics
Upcoming discussion: Microbial symbioses, including:
Bacteria–Fungi interactions
Bacteria–plant interactions
Fungi–plant interactions
Emphasis on ecological relationships and their relevance to health, agriculture, and ecosystems.
Quick Reference: Key Terms and Concepts
Protists: a broad group of unicellular eukaryotes (excluding yeasts/algae in some contexts).
Mycelium: filamentous network of fungal hyphae.
Hyphae: thread-like filaments forming fungal structures.
Peptidoglycan: mesh-like polymer in bacterial cell walls; synthesis involves transglycosylation and transpeptidation.
Capsid: protein shell encasing the viral genome; built from capsomeres.
Nucleocapsid: genome + capsid complex.
Enveloped virus: virion with a lipid membrane surrounding the capsid.
Baltimore classification: system grouping viruses by genome type and replication strategy (seven classes).
Lytic cycle: phage replication culminating in host cell lysis and virion release.
Lysogenic cycle: phage genome persists within host, can later re-enter lytic cycle.
CRISPR-Cas: bacterial adaptive immune system against phages; basis for powerful biotechnologies.
Notable Numerical and Structural Details (Examples)
Virus sizes (examples):
Poliovirus: 30 ext{ nm}
Smallpox virus: 200 ext{ nm} imes 300 ext{ nm}
Bacteriophage T4: 50 ext{ nm} imes 225 ext{ nm}
Bacteriophage MS2: 24 ext{ nm}
TMV: 15 ext{ nm} imes 300 ext{ nm}
Cellular and bacterial sizes:
E. coli: 1{,}000 ext{ nm} imes 3{,}000 ext{ nm}
Red blood cell: 10{,}000 ext{ nm} in diameter
Nucleic acid types and formats are summarized in the Baltimore classes (I–VII) with respective genome types and strategies as described above.