17/4 Lecture Notes on Proteins and Membrane Transport
Introduction to Proteins
Today's focus is on proteins, particularly an ADP ATP transporter.
An average human cell can express around 20,000 different proteins, with about 10,000 present depending on cell type.
Approximately 2,000 proteins are associated with membranes, facilitating communication outside the cell or organelle.
Organelles separate important pathways and processes within the cell.
Membrane proteins:
Receptors: Recognize signals (e.g., hormones like glucagon receptor).
Channels: Allow ions to pass in and out.
Transporters: Require some input for controlled movement of molecules.
Levels of Protein Structure
Primary Structure: The amino acid sequence.
Secondary Structure: Organization of amino acids into structures like alpha helices and beta sheets. (chain organisation, domains)
Tertiary Structure: How secondary structure elements combine to form a functional protein. (domain organisation, ordered peptide)
Quaternary Structure: Association of multiple protein subunits for function. (interaction of multiple structured peptides)
Example: Aldolase, which functions as a tetramer (four subunits).
Stabilization of Protein Structures
Primary Structure:
Amino acids are linked by peptide bonds (carbon and nitrogen linkage).
Peptide bond is stable and has a planar arrangement, important for protein folding.
Condensation reaction: water is released.
Peptide bonds broken by proteases, not by detergents used in aldolase prac.
Acid hydrolysis required to break these bonds under harsh conditions
Amino Acids and Their Characteristics
20 different amino acids with varying R groups.
Polar amino acids: hydrophilic and interact with water.
Polar uncharged.
Polar charged: Positively and negatively charged R groups.
Hydrophobic non polar amino acids: avoid interaction with water.
The arrangement of these amino acids determines protein folding.
Weak Interactions Driving Protein Folding
Protein folding is driven by a combination of weak interactions:
Hydrogen bonds: interaction between electronegative atom (oxygen) and electropositive hydrogen.
Electrostatic interactions: interactions between oppositely charged amino acid side chains.
Van der Waals forces: temporal fluctuations in electron positions around atoms.
Hydrophobic effect: hydrophobic amino acids shield themselves from water and are buried inside the protein. (fats)
Secondary Structures
Alpha Helices:
Hydrogen bonding between backbone residues.
Side chains project radially from the helix, not involved in stabilisation of structure.
Held together by hydrogen bonds between carbonyl and groups.
One turn of the alpha helix requires residues.
Beta Sheets:
Composed of beta strands. Strands are to residues long.
Hydrogen bonding between peptide backbones.
R groups project differently depending on whether it's a parallel or anti-parallel beta sheet.
Loops:
Connect secondary structures.
Often have important functional roles
Cytochrome C Oxidase Example
Membrane protein with helices situated in the lipid membrane.
Loops and functional groups in the aqueous region.
Hydrophobic amino acids are associated with helices that interact with the hydrophobic lipid regions.
Tertiary Structure
Arrangement of secondary structure elements.
Driven by side chain interactions.
Conserved amino acids in regions important for function, especially in enzymes.
Proteins with almost no primary amino acid level sequence identity can have identical folds but not necessarily identical functions.
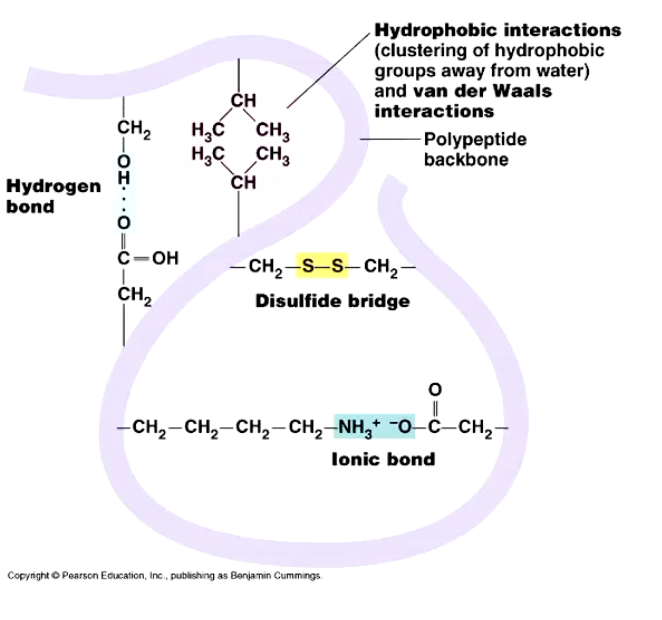
Examples of interactions: Hydrophobic interactions, hydrogen bonding, and ionic interactions.
Disulfide bridges: Strong bonds formed via oxidation, often associated with extracellular proteins in harsh environments; broken by reducing agents
Energetically stable molecule arranged to satisfy interactions.
Structural Motifs
Proteins grouped by how structures come together (e.g., helix-loop-helix).
interacting helices are arranged so that side-chains of one helix slot into the groove of a neighbour
Beta-barrel structures:
Common in membrane proteins and bacterial toxins.
Can form pores in cells.
Quaternary Structure
How tertiary structure units come together.
Homo-oligomers: Multiple subunits of the same protein (dimers, trimers, tetramers).
Hetero-oligomers: Multiple subunits of different proteins (e.g., F1ATPase).
Function of Membrane Proteins
Capacity to perceive signals from outside the cell.
Translate signals to change pathways inside the cell.
Transporters facilitate movement of molecules.
ATP ADP Translocase
Crucial protein for ATP production.
We only have around grams of ATP in our body, and so what we need to do is we need to generate around about body weight in ATP every day to survive.
Most ATP production occurs in the mitochondria (some in cytosol).
ATP ADP Translocase Abundant protein (around % of inner mitochondrial membrane protein).
Facilitates stoichiometric (one-to-one) transport of ADP into the mitochondrial matrix and ATP out.
Electrogenic transport: Influenced by charge difference across the membrane.
Intermembrane space: positively charged due to protons.
Matrix: negatively charged.
Favors ATP transport out (more negatively charged due to additional phosphate).
Phosphate is also transported in, important for ADP to ATP synthesis.
Structure of ATP ADP Translocase
Six transmembrane helices (H1-H6) that embed in the membrane.
Hydrophobic amino acids align with the lipid part of the membrane, while hydrophilic amino acids enable association with ATP and ADP.
Two different conformations:
Open to the mitochondrial matrix to accept ATP binding.
Open to the intermembrane space to accept ADP binding.
Cavity lining (hollow space/pockets) is hydrophilic.
Lipid lining residues are hydrophobic.
Charged residues (lysine, arginine) at the entrance of the cavity bind ATP (start bringing atp into cavity of protein).
Aromatic residues (tyrosine) direct the sugar region.
ATP binding forces a conformational change:
Conformational change releases ATP.
Transporter then accepts ADP for transport back across.
Role of Cardiolipins
Cardiolipins are crucial for the activity of the protein.
Disorders affecting cardiolipin production lead to muscle wastage and low energy production.
Cardiolipins stabilize the structure in the membrane and facilitate conformational change.
Prevent misfolding or aggregation of the protein.
Additional points:
ATP, ADP are polar molecules, so they need a transporter to go across, and that enables us to control the transport of these really important molecules.
Lipid membranes have hydrophobic regions.
Key points to review for your exam:
Protein Structure Levels: Primary, secondary, tertiary, and quaternary structures, and what each entails.
Stabilization of Protein Structures: Focus on peptide bonds, their characteristics, and how they're broken.
Amino Acids: The different types (polar, hydrophobic) and their roles in protein folding.
Weak Interactions: Hydrogen bonds, electrostatic interactions, Van der Waals forces, and the hydrophobic effect.
Secondary Structures: Alpha helices, beta sheets, and loops—their composition and importance.
Cytochrome C Oxidase: Understand its structure as a membrane protein and the arrangement of amino acids.
Tertiary Structure: How secondary structures arrange and the types of interactions involved. Pay special attention to disulfide bridges